Back to the future—breast cancer reprises pathways found in fetal cells
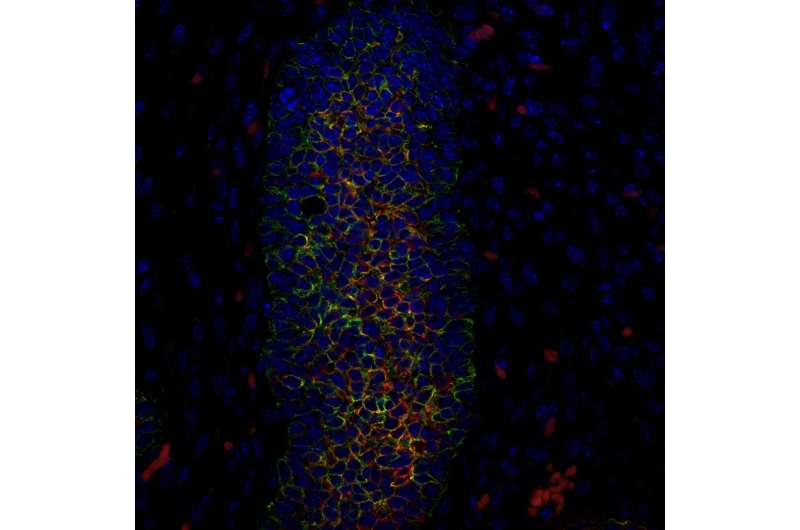
Using just a microscope, Italian surgeon Francesco Durante was struck by the similarities between cells in the most malignant cancers and the embryonic cells of the organ in which the cancer originated.
More than a century later, scientists at the Salk Institute have uncovered a reason for the uncanny likeness: cells in human basal-like breast cancers share features with the embryonic mammary (breast) stem cells that are the progenitors of all cell types in the mammary gland (of a mouse). The insights leading to this conclusion are published in the journal Cell Reports on August 7, 2018.
"Durante was prescient," says Professor Geoffrey Wahl, holder of the Daniel and Martina Lewis Chair and senior author of the work. "He anticipated the relatedness of cells in the embryo to those in malignant cancers—and that dormant cancer cells could be 'reawakened' by exposure to 'persistent irritations' that we now recognize as inflammation. We can use the insights gained from our work to develop better diagnostic and treatment strategies."
For example, human breast cancers share some peculiar metabolic features with early embryonic mammary stem cells, which may be possible to target therapeutically. Additionally, proteins specifically expressed in the embryonic cells that are also expressed in the cancers may be used to develop new diagnostic agents or tools for immune therapies.
Cancer has been called a "caricature of development," reprising features of the embryonic stem cell state for their own perverse purposes. So Wahl and his research group at Salk, along with investigator Benjamin Spike of the Huntsman Cancer Institute at the University of Utah, used cutting-edge techniques to generate an atlas of the genes expressed in each breast cell from very early in development until adulthood, a process that required an analysis of many thousands of cells. They used this "single-cell-transcriptome atlas" to compare genes expressed in human breast cancers. This led to an understanding of how the stem cells of the breast arise in early development and how they turn into the two different types of cells that comprise the mature gland.

"There has been intense interest in determining how rare cells in tumors can fuel tumor growth and resistance to therapies," says Spike, who is an assistant professor of oncological sciences at the University of Utah and the paper's co-corresponding author. "Much of the molecular machinery they use to do this appears to be co-opted and corrupted from stem cells and progenitors that used this machinery to build the normal tissue during development. Our study provides an atlas of the responsible genes that can be tested for their potential as therapeutic targets."
"This work shows the diversity of ways that cells can enter the stem state, which is characterized by their plasticity, or developmental flexibility," adds first author Rajshekhar Giraddi, a Salk research associate in Wahl's lab. "This suggests that cancer cells may gain their plasticity by many strategies, similar to those we are discovering in normal development."
This developmental plasticity likely explains how the cells within a single tumor can appear so different from one another and likely underlies the uncanny ability of malignant cancer cells to become resistant to most therapies.
Now, armed with this knowledge of the genetic signatures of different cell states, the lab is developing new ways of looking at the reprogramming of adult cells into states associated with cancer.
"What would be great is if we can figure out how to prevent the reprogramming of cancer cells to become so developmentally plastic." says Wahl. "This plasticity will likely preclude development of a single 'magic bullet' to treat cancer. Rather, cancers are very adaptive diseases, requiring attacking them from multiple directions."
More information: Rajshekhar R. Giraddi et al, Single-Cell Transcriptomes Distinguish Stem Cell State Changes and Lineage Specification Programs in Early Mammary Gland Development, Cell Reports (2018). DOI: 10.1016/j.celrep.2018.07.025