Genomic study finds a new role for microRNAs as predictors of Crohn's disease progression
Crohn's disease is a lifelong condition characterized by a fluctuating course of gastro-intestinal inflammation with repeated flares and remissions. Any part of the alimentary tract from the mouth to the anus can be affected resulting in diverse symptoms including abdominal pain, watery diarrhea, and hematochezia. Furthermore, persistent inflammation can result in complications, such as a narrowing of the large intestine called strictures or perforations in the intestinal wall, both of which often require surgical treatment and severely compromise the quality of life of Crohn's patients. The incidence of Crohn's disease has increased throughout the world over the last 50 years for all ages, indicating its emergence as a global disease.
Currently available drugs for Crohn's disease also increase the potential risk of serious side effects including opportunistic infection and cancer development. Because disease course widely varies among patients and no "one size fits all" treatment exists, determining which patients are at high risk for poor clinical outcome is a critical problem in the management of Crohn's disease.
Now a new study conducted in adult and pediatric patients with Crohn's disease, led by UNC School of Medicine researchers and published in the Journal of Clinical Investigation (JCI) Insight, has found that a set of biomolecules known as microRNAs, specifically microRNA-31 (miR-31), can help predict which patients with Crohn's disease are at higher risk for the development of severe problems that may require surgical removal of the large intestine.
"For such a clinically heterogenous disease, this kind of molecular phenotyping is a major step towards personalization of medical therapy, said co-senior author Shehzad Z. Sheikh, MD, Ph.D., associate professor of medicine and genetics. "These results add to a series of papers from our group where we combine genomic technologies with rigorous validation in patient-derived, disease-relevant cell systems to develop molecular markers with prognostic utility."
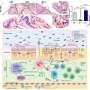
A key validation step in this research involved the generation of organoids or 'mini-guts' that architecturally and physiologically resemble the human intestine. Sheikh and colleagues derived organoids from Crohn's patients to preserve the molecular defects observed in the patient tissue.
"This innovative system can serve as a personalized testing platform to screen therapeutic agents before administering them to the patient," Sheikh said."
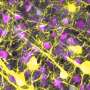
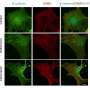
In the study, small RNA-sequencing was performed on adult colon tissue from 18 adults with Crohn's disease and 12 controls. Small RNA-sequencing was also performed on colon tissue from 76 children who were newly diagnosed with Crohn's but had not received any treatment yet, and 51 controls. In addition to these whole tissue assays, colonic epithelial cells and immune cells were isolated from colonic tissues and miR-31 expression was measured.
These analyses found that in adults, low colonic miR-31 expression at the time of surgery was associated with worse disease outcomes, requiring an end ileostomy and later recurrence of disease. In children, low colonic miR-31 expression at the time of diagnosis was found to be associated with future development of strictures that required surgery.
"In recent years there has been great success in deriving molecular signatures that classify several cancers into subtypes, including studies performed here at UNC," said co-senior author Terry Furey, Ph.D., associate professor of genetics and biology and a member of the UNC Lineberger Comprehensive Cancer Center. "These subtypes, then, have been shown to be useful in determining cancer progression and response to specific therapies. Our long-term goal, extending the work in this study, is to uncover molecular subtypes of Crohn's disease to not only increase our understanding of the root causes of the disease and the vast clinical heterogeneity, but also to more strategically use current therapies and provide the basis for new therapies that specifically target these subtypes."
The third co-corresponding author is Praveen Sethupathy, Ph.D., associate professor of biomedical sciences at Cornell University, who established a collaboration with Drs. Sheikh and Furey while he was a faculty member at UNC from 2011-2017. "While microRNAs as prognostic indicators of disease development has emerged as an exciting concept, there have been very few clinically actionable examples to date. We are excited about our findings because, while it is just the beginning and more work remains to be done, it opens the possibility of using microRNAs to improve clinical trial designs for Crohn's disease and developing more personalized therapeutic strategies for patients. Also, our study emphasizes the importance of epithelial biology in Crohn's disease, which merits deeper investigation to delineate mechanisms that underlie the etiology of different disease subtypes," Sethupathy said.
Benjamin P. Keith, a graduate student in the Sheikh Lab, is first author of the study. He highlighted the importance of the methods the researchers used in the clinic to preserve patient tissue samples.
"Preserving tissue in this way usually results in heavily degraded mRNA," Keith said. "But our ability to accurately detect microRNA expression that was not degraded in fixed samples opens up future studies to a wealth of patient samples from across the country to further validate microRNA-31 and a suite of other microRNAs as prognostic markers of Crohn's disease," Keith said.