Microbiome profiling reveals associations with ulcerative colitis severity, treatment
A study of gut microbes from more than 400 children points to how the microbiome behaves in this inflammatory bowel disease.
The gastrointestinal tract hosts a complicated set of relationships involving a variety of host cells (e.g., epithelial cells, immune cells) and the array of microbes that make their home inside the gut (collectively known as the gut microbiome). In ulcerative colitis (UC) and Crohn's disease, those relationships are interrupted by mechanisms that researchers do not yet fully grasp, but in which the microbiome may play a critical role.
To better understand where microbes fit in UC progression and treatment, a multicenter team led by computational scientist Melanie Schirmer and core institute member Ramnik Xavier of the Broad's Infectious Disease and Microbiome Program (IDMP) profiled the gut microbiomes of 405 children with UC for a year after diagnosis.
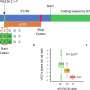
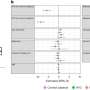
By bringing comprehensive health records together with 16s rRNA amplicon sequencing—a sequencing approach that gives a snapshot of all microbes within a sample—of patient stool and gut biopsies, the team found clear patterns in microbiome composition that tracked over time with disease severity and treatment response. They reported their findings in Cell Host & Microbe.
"Most of the microbiome profiling work related to ulcerative colitis has focused on patients on treatment," said Xavier, who co-directs both the IDMP and the Center for Microbiome Informatics and Therapeutics (CMIT) at the Massachusetts Institute of Technology. He also directs the Center for Computational and Integrative Biology at Massachusetts General Hospital. "This study has helped reveal features of the treatment-naïve microbiome in pediatric UC, and identify links between the microbiome and refractory disease."
All told, the researchers linked the abundance of 50 microbial organisms with disease severity at diagnosis. They also found that changes in microbial abundances over time associated with whether or not a child's disease responded to standard therapies. This included the surprising finding that microbiomes from children with severe UC were unusually rich in bacteria normally found in the mouth, which exacerbate inflammation in UC patients.
The team also noted that the levels of certain microbe-targeting antibodies changed in UC patients. These changes also tracked with disease severity, and suggested that interactions between the immune system and certain gut microbes may influence whether a child's disease gets worse or improves over time.
The authors pointed out that they cannot say whether the patterns they found help cause UC. However, their findings suggest that microbiomes might serve as therapeutic resources for UC treatment, as biomarkers for forecasting a patient's clinical course, and for planning and monitoring care.
Schirmer and Xavier's team carried out their analysis under the auspices of PROTECT, a National Institutes of Health-supported study aimed at providing a better understanding of how children newly diagnosed with UC respond to standard initial therapies.
More information: Melanie Schirmer et al. Compositional and Temporal Changes in the Gut Microbiome of Pediatric Ulcerative Colitis Patients Are Linked to Disease Course, Cell Host & Microbe (2018). DOI: 10.1016/j.chom.2018.09.009