Scientists grow functioning human neural networks in 3-D from stem cells
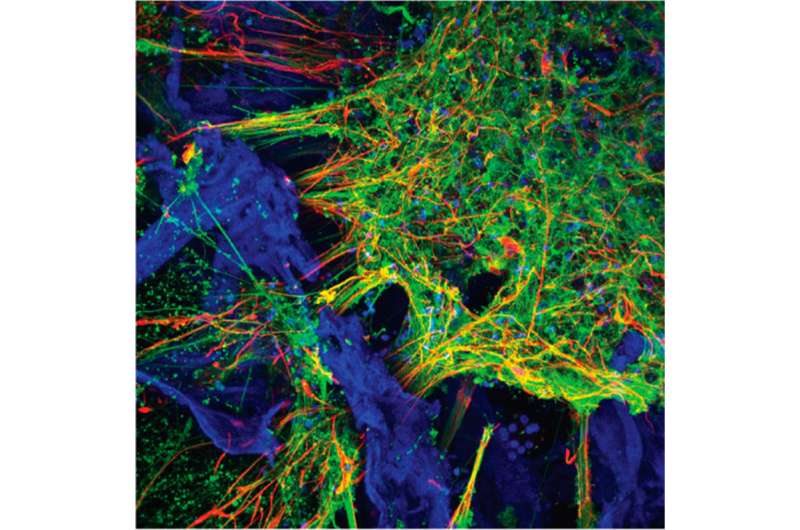
A team of Tufts University-led researchers has developed three-dimensional (3-D) human tissue culture models for the central nervous system that mimic structural and functional features of the brain and demonstrate neural activity sustained over a period of many months. With the ability to populate a 3-D matrix of silk protein and collagen with cells from patients with Alzheimer's disease, Parkinson's disease, and other conditions, the tissue models allow for the exploration of cell interactions, disease progression and response to treatment. The development and characterization of the models are reported today in ACS Biomaterials Science & Engineering, a journal of the American Chemical Society.
The new 3-D brain tissue models overcome a key challenge of previous models -the availability of human source neurons. This is due to the fact that neurological tissues are rarely removed from healthy patients and are usually only available post-mortem from diseased patients. The 3-D tissue models are instead populated with human induced pluripotent stem cells (iPSCs) that can be derived from many sources, including patient skin. The iPSCs are generated by turning back the clock on cell development to their embryonic-like precursors. They can then be dialed forward again to any cell type, including neurons.
The 3-D brain tissue models were the result of a collaborative effort between engineering and the medical sciences and included researchers from Tufts University School of Engineering, Tufts University School of Medicine, the Sackler School of Graduate Biomedical Sciences at Tufts, and the Jackson Laboratory.
"We found the right conditions to get the iPSCs to differentiate into a number of different neural subtypes, as well as astrocytes that support the growing neural networks," said David L. Kaplan, Ph.D., Stern Family Professor of Engineering, chair of the Department of Biomedical Engineering at Tufts' School of Engineering and program faculty member at the Sackler School of Graduate Biomedical Sciences at Tufts. "The silk-collagen scaffolds provide the right environment to produce cells with the genetic signatures and electrical signaling found in native neuronal tissues."
Compared to growing and culturing cells in two dimensions, the three-dimensional matrix yields a significantly more complete mix of cells found in neural tissue, with the appropriate morphology and expression of receptors and neurotransmitters, according to the paper.
Others have used iPSCs to create brain-like organoids, which are small dense spherical structures useful for understanding brain development and function, but can still make it difficult to tease out what individual cells are doing in real time. Also, cells in the center of the organoids may not receive enough oxygen or nutrients to function in a native state. The porous structure of the 3-D tissue cultures described in this study provides ample oxygenation, access for nutrients and measurement of cellular properties. A clear window in the center of each 3-D matrix enables researchers to visualize the growth, organization and behavior of individual cells.
"The growth of neural networks is sustained and very consistent in the 3-D tissue models, whether we use cells from healthy individuals or cells from patients with Alzheimer's or Parkinson's disease," said William Cantley, Ph.D., 2018 graduate of the Cell, Molecular & Developmental Biology program at the Sackler School of Graduate Biomedical Sciences at Tufts and first author of the study, which was completed as part of his Ph.D. dissertation. "That gives us a reliable platform to study different disease conditions and the ability to observe what happens to the cells over the long term."
The researchers are looking ahead to take greater advantage of the 3-D tissue models with advanced imaging techniques, and the addition of other cell types, such as microglia and endothelial cells, to create a more complete model of the brain environment and the complex interactions that are involved in signaling, learning and plasticity, and degeneration.
More information: William L. Cantley et al, Functional and Sustainable 3D Human Neural Network Models from Pluripotent Stem Cells, ACS Biomaterials Science & Engineering (2018). DOI: 10.1021/acsbiomaterials.8b00622