DF-PGT, now possible through massive sequencing techniques
A research team from the Universitat Autònoma de Barcelona (UAB), in collaboration with the Blood and Tissue Bank of Catalonia, has implemented a massive sequencing platform for preimplantation genetic testing (PGT) for the first time in history.
The work adapts the TruSight One (TSO) platform, one of today's most complete genetic panels, to Double Factor Preimplantation Genetic Testing (DF-PGT). The study, published in PLoS ONE, was led by Joaquima Navarro and Jordi Benet, researchers from the UAB Department of Cellular Biology, Physiology and Immunology, and included the collaboration of the team led by Francisco Vidal from the Blood and Tissue Bank of Catalonia.
"We successfully managed to implement an innovative, promising and universal strategy, prepared for a simultaneous diagnosis of genetic mutations and chromosomal alterations within embryos obtained by in vitro fertilisation (IVF), of benefit to the DF-PGT candidate families with mutations causing diseases included in the TSO platform. In addition, it only requires a single laboratory experiment, without the need to prepare the diagnosis methodology. This substantially speeds up the study process and the availability of the results of the family's single gene disorders. Until now, there was a need to prepare the specific procedures before conducting the diagnosis for each of the mutations," Joaquima Navarro says.
The new tool will make it possible to diagnose mutations both directly and indirectly, which increases the degree of diagnostic accuracy. At the same time, it allows for chromosomal characterisation within embryos for the totality of all 23 human chromosome pairs, and detects whether the embryo is aneuploid, having an abnormal number of chromosomes, or euploid, with the correct number and therefore viable and with greater chances of implantation.
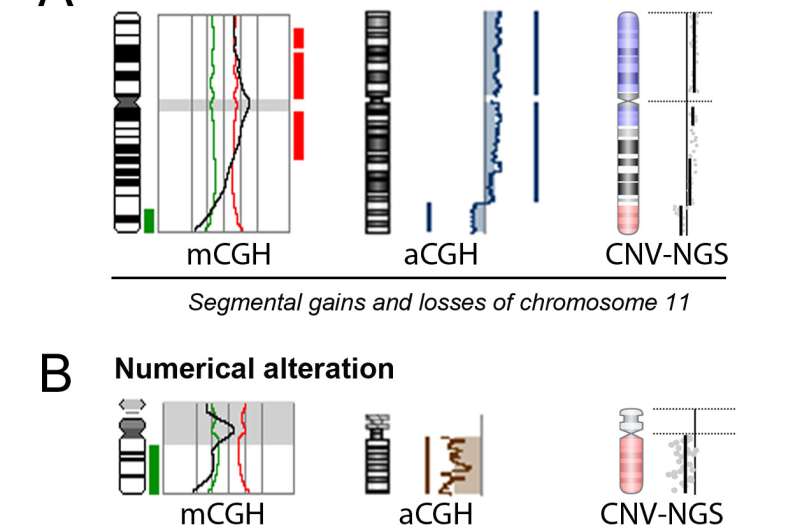
The team led by Professor Navarro developed the DF-PGT strategy in 2009. A pioneering technique successfully applied on numerous occasions since then, it consists of analysing within one IVF cycle the specific genetic mutations causing hereditary diseases, as well as a complete embryonic chromosomal endowment (cytogenetics) through a comparative genomic hybridisation technique. This allows identifying and selecting embryos free of diseases and chromosomal defects that would hinder their development.
In 2009, 2013 and 2015, this group was the first ever to use another DF-PGT strategy to select healthy embryos and help families conceive healthy offspring, including twins free of Von Hippel-Lindau syndrome, twins free of Lynch syndrome, and two other healthy babies from two families at risk from sickle cell disease and cystic fibrosis, respectively. On those occasions, the scientists first had to prepare the diagnostic methods of the mutations responsible for the specific genetic disease, whereas in the current strategy proposed, no specific preparation is necessary.
Overcoming DNA Limitations in Embryonic Cells
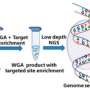
Next-generation sequencing (NGS) techniques represent a giant leap forward in the quality of genetic analysis procedures,permitting the study of millions of DNA sequences massively and simultaneously in one experiment. These powerful tools are being used for the characterisation of blood and tissue samples in which the amount of DNA is not a restricting factor.
"The proposed methodology overcomes the limitations existing until now. It was developed to be applied to samples of only six to eight blastocyst trophoderm cells with the scheduling of a frozen embryo transfer cycle, in the case of the results indicating an absence of family diseases and embryo," explains Joaquima Navarro.
Before implementing the new platform, there was a need to determine which of the four most commonly used DNA amplification systems was most suitable for the adequate identification of mutations. Researchers were thus also the first ever to conduct a chromosomal characterisation with the Nexus computational biology programme by using the TSO database.
"The tool is also of interest in PGTs with risk of chromosomal alterations due to advanced maternal age, alterations found in the father's chromosomes, as well as in cases of repeated miscarriages. Also in cases of the oocytes of young donors, since a certain risk of aneuploidy has also been described," the UAB researcher says.
This new methodology will soon be available for use with the TSO panel or by applying it to new, even more complete panels, such as the ones capable of analysing the whole exome or whole genome sequencing. In the middle term, it can be applied simultaneously to many samples, thereby reducing its cost.
More information: Javier del Rey et al, Novel Double Factor PGT strategy analyzing blastocyst stage embryos in a single NGS procedure, PLOS ONE (2018). DOI: 10.1371/journal.pone.0205692