New approaches to study the genetics of autism spectrum disorder may lead to new therapies
Canadian neuroscientists are using novel experimental approaches to understand autism spectrum disorder, from studying multiple variation in a single gene to the investigation of networks of interacting genes to find new treatments for the disorder.
Autism spectrum disorder (ASD) affects more than 1% of children, yet most cases are of unknown or poorly defined genetic origin. It is highly variable disorder, both in its presentation and in its genetics—hundreds of risk genes have been identified. One key to understanding and ultimately treating ASD is to identify common molecular mechanisms underlying this genetically heterogeneous disorder. Four Canadian researchers presented the results of unique approaches to understand ASD at the 14th Canadian Neuroscience Meeting in Toronto, on May 24, 2019.
One common feature of autism is a shift in the ratio of excitation (or activation) and Inhibition (or inactivation) of neurons in animal models of ASD. Mutations that cause too much excitation of neurons result in autistic-like behaviour, and paradoxically, so do mutations that cause too much inhibition. Precise control of the Excitation to Inhibition ratio is therefore viewed as a key to regulate social behaviour. Dr. Melanie Woodin, at the University of Toronto, investigated a protein that is critically important for neuronal inhibition, called KCC2. When KCC2 fails to work, inhibitory neurotransmission (through a neurotransmitter called GABA) switches to being excitatory. Breakdown of GABA inhibition is a hallmark of abnormal brain activity in conditions such as epilepsy, pain and some forms of autism.
Regulation of KCC2 therefore appears as a valid target for treatment of ASD. Dr. Woodin's team has identified the first comprehensive list of proteins that interact and modify the action of KCC2. Their work has shown that one protein, called Pacsin1, interacts with KCC2 and can regulate its abundance and localisation. These results suggest that manipulating KCC2 interacting proteins could be an efficient technique to regulate KCC2 in a neuron specific manner.
More than a thousand mutations and other forms of genetic variation affecting several hundred genes have been linked to ASD. Given this large number, analyzing each gene on its own is not a feasible approach. To make sense of this data, one approach is to determine whether multiple risk genes function in common signaling pathways, which act as "hubs" where risk genes converge. To identify such hubs or networks, Dr. Karun Singh, from McMaster's University studies proteins in mouse models of ASD, but also in cells taken from patients and induced to grow in petri dishes, called induce pluripotent stem cells or iPSCs. By looking at how the proteins taken from cells carrying ASD associated mutations interact, his team has been able to identify specific signaling pathways affected by ASD. Targeting of these networks may lead to new therapies for ASD.
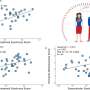
Dr. Catharine Rankin, from the University of British Columbia, presented data obtained by analyzing ASD-associated genes in a much simpler species, the nematode worm C. elegans. Her team tested 87 different strains of worms, each carrying a mutation in genes similar to those found in ASD-associated genes. Analysis of the morphology, locomotion, sensitivity and habituation, which is the simplest form of learning, in these worms by an automated system revealed certain genes that caused strikingly similar effect on the worms. Further analysis revealed these similarities resulted from previously undescribed interactions between the affected genes.
A great advantage of studying ASD genes in nematode worm is the possibility to easily edit genes and study the effects of these modifications on the worm by automated systems. This provides a means to analyse a large range of genes, thereby revealing unique and/or shared functions. Candidate drugs can also be tested for their ability to rescue the deficit associated with different gene modifications. Furthermore, Dr. Rankin demonstrated the feasibility of using the gene editing system based on CRISPR-Cas9 to specifically insert or remove ASD-related genes at specific times, to study their role in development.
The final speaker in this session was Dr. Kurt Haas, from the University of British Columbia, who discussed the role of a gene called PTEN. Mutations in PTEN have been strongly linked to both cancer and ASD, yet the mechanisms through which this occurs was unclear. Dr. Haas reported on the results obtained by 7 laboratories at that institution who collaborated to test 105 variants of PTEN, in yeast, fly, worm, rat, and human cell lines, to understand the impact of different mutations in this gene in a wide diversity of cellular environments. This analysis allowed the researchers to determine the specific impact ASD associated mutations on various protein functions, with high confidence.
By using a range of different approaches, Drs. Woodin, Singh, Rankin and Haas have increased our understanding of the genetic underpinnings of autism spectrum disorder. These studies pave the way to the identification of new potential therapeutic targets to treat this disorder.