After GWAS studies, how to narrow the search for genes?

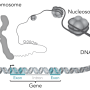
Genome-wide association studies (GWAS) look at large populations to find genes that contribute to common, multi-gene traits like height or obesity. These comprehensive investigations frequently turn up large numbers of tiny genetic variations that show up more often in people who are tall, obese, etc. But this association doesn't mean the variant actually helps cause the trait; it could just be going along for the ride.
So which genes should scientists prioritize for further investigation? Numerous computational algorithms are available to help distill GWAS results, each using different criteria and assumptions. But it's been hard to know which one to pick.
Most methods used to evaluate such algorithms can bias investigators toward genes that are already well-characterized, steering them away from opportunities to discover something truly new. Other methods require access to independent reference data that aren't always readily available.
"We have different prioritization algorithms, but we don't actually know how to decide which one is best," says Rebecca Fine, a PhD candidate at Harvard Medical School who has been working on this problem. "We didn't want to have to rely on a previous 'gold standard' or bring in anything other than the original GWAS data."
Fine and Joel Hirschhorn, MD, PhD, chief of endocrinology at Boston Children's Hospital, have developed what they believe is an effective, unbiased method called Benchmarker, described in the American Journal of Human Genetics earlier this month.
Borrowing from machine learning
Borrowing the machine-learning concept of "cross-validation," Benchmarker enables investigators to use the GWAS data itself as its own control. The idea is to take the GWAS dataset and single out one chromosome. The algorithm being benchmarked then uses the data from the remaining 21 chromosomes (all but X and Y) to make predictions about what genes on the single chromosome are most likely to contribute to the trait being investigated. As this process is repeated for each chromosome in turn, the genes that the algorithm has flagged are pooled. The algorithm is then validated by comparing this group of prioritized genes with the original GWAS results.
"You train the algorithm on the GWAS with one chromosome withheld, then go back to that chromosome and ask whether those genes were actually associated with a strong p-value in the original GWAS results," explains Fine. "While these p-values don't represent the exact 'right answers,' they do tell you roughly where some true genetic associations are. The end product is an evaluation of how each algorithm performed."
Benchmarking Benchmarker
Putting this approach through its paces for 20 separate traits, Fine, Hirschhorn and colleagues conclude that combining multiple strategies often gives the best results. They also found evidence that certain algorithms perform best when looking for genes for certain traits.
"We expect that many more algorithms will be developed to answer the key next question after GWAS: which genes and variants are causally related to human traits and diseases," says Hirschhorn. "The Benchmarker approach can be a great help as an unbiased way to figure out which algorithms to use to answer this question."
More information: Rebecca S. Fine et al, Benchmarker: An Unbiased, Association-Data-Driven Strategy to Evaluate Gene Prioritization Algorithms, The American Journal of Human Genetics (2019). DOI: 10.1016/j.ajhg.2019.03.027