March 13, 2019 report
Researchers identify causal variants in blood cells and tie them with genetic mechanisms
A team of researchers affiliated with multiple institutions in and around the Boston area in the U.S. and one in Finland has identified hundreds of possible causal variants associated with blood cell traits, and tied them to important blood-related mechanisms.
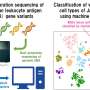
The team sought to learn more about genetic variants in blood cells that are responsible for blood mechanisms such as which genes are responsible for regulating the number of white blood cells, and which regulate the optimal number of white blood cells for an individual. To that end, they conducted a genome-wide association study (GWAS) by accessing data in the UK Biobank, which provided information regarding blood cell characteristics such as hemoglobin levels, white cell count and platelet count. The data represent health information for approximately 115,000 people in the United Kingdom. The traits the team focused on centered around those involved in hemopoiesis (production of blood cells and platelets), which occurs in bone marrow.
The team used fine mapping to find possible candidates as they examined 2,000 3-Mb-sized genetic regions that had a genome-wide association. They were able to identify 38,654 variants that their math suggested had a better than 1 percent probability of being causal. They also found that those variants with a posterior probability greater than 0.75 also had a minor allele frequency that was more than 5 percent.
The researchers then filtered out those variants that were nonsynonymous and those that were identified as part of loss-of-function coding in 230 genes that were linked to the mechanisms they were focusing on, such as monocytes, lymphocytes, red blood cells and platelets. This allowed them to identify the genes that could be associated with particular mechanisms, such as traits of red blood cells that were connected to iron homeostasis, or traits of platelets that could be tied to the process of coagulation. They also noted that more than 170 variants they identified were pleiotropic.
As part of their efforts, the researchers also developed a new way to enrich the results of fine-mapped variants, which they called genetic-chromVAR.
More information: Jacob C. Ulirsch et al. Interrogation of human hematopoiesis at single-cell and single-variant resolution, Nature Genetics (2019). DOI: 10.1038/s41588-019-0362-6
© 2019 Science X Network