Novel tool used to mine clinical data and identify causative gene in childhood epilepsy

A team of researchers at Children's Hospital of Philadelphia (CHOP) affiliated with the CHOP Epilepsy Neurogenetics Initiative (ENGIN) discovered a new gene associated with severe childhood epilepsy using a novel computational approach. The team systematically compared phenotypes, or clinical data, of patients with severe childhood epilepsies through a novel analysis strategy and looked for common genetic causes in patients who had similar clinical presentations.
This is the first time that such an analysis of clinical data has been used to identify novel genetic causes of neurological disorders, and this new computational method has the potential to help patients with a variety of complex and difficult-to-diagnose conditions. The findings were published today in the American Journal of Human Genetics.
"Genetic data are incredibly valuable, but when we do whole exome sequencing, this is really only half of the story," said Ingo Helbig, MD, a pediatric neurologist in the Division of Neurology at CHOP who directs the Genomics and Datascience Core of the CHOP Epilepsy Neurogenetics Initiative. "Genetic epilepsies can present with a wide range of symptoms, and what we really want to understand is which medications work and how we can improve outcome. Genetic testing alone does not give us this information. However, when we merge genetic information with large-scale clinical data, the combination can be very powerful."
Developmental and epileptic encephalopathies are severe heterogeneous brain disorders that often have a genetic cause. However, the genetic basis for these disorders remains unknown in a large portion of affected patients. These disorders can cause aggressive seizures, cognitive and neurological deficits and, in some cases, early death. Many patients do not respond to current treatment options, and identifying a causative gene is often the first step of improving treatment.
Over the last decade, technological advances have made it possible for genetic testing to be performed on a large scale, yielding large sets of genomic data that have enabled researchers to pinpoint a number of important genetic mutations in childhood epilepsies. However, the patient's phenotype—a set of observable characteristics such as type of seizures or developmental disabilities—has historically not been collected in the same standardized manner as genetic data. While new gene sequencing technologies can generate genomic data more thoroughly and quickly, clinical data must be entered by hand, which results in a "phenotypic bottleneck" where clinical data cannot be processed at the same level as genetic data.
To address this discrepancy, other researchers previously developed the Human Phenotype Ontology (HPO), a catalogue that provides a standardized format to characterize a patient's phenotypic features, including neurological findings, and allows for clinical data to be used at a similar level as genetic data.
Since many severe childhood epilepsies have very complex clinical findings, the study team believed that analyzing clinical data in the HPO data through computational methods would be an effective way to identify patients with similar symptoms that may not have been obvious. By combining this phenotypic information with a patient's whole exome sequencing data, researchers wanted to see if similarities in clinical features might reveal information about the genetic basis of the disorder that might otherwise go undetected.
"The main limitation in the past was the lack of large amounts of clinical information in a format that can be analyzed systematically through our informatics approaches," Helbig said. "In our study, we built the computational algorithms to leverage clinical data. We then used these tools to find the genetic cause for a patient's epilepsy."
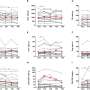
Helbig and colleagues, including collaborators from the EuroEPINOMICS-RES Consortium and the U.S.-based Genomics Research and Innovation Network, analyzed whole exome sequencing data and clinical data transformed to HPO format for computational analysis in 314 patients with severe epilepsies. The team found a variant in the AP2M1 gene in two individuals with similar phenotypes in their study. Using this information, they looked for variants in AP2M1 in a second cohort of 2,310 individuals and found two additional patients with similar clinical features, including neurodevelopmental disorders and generalized epilepsy.
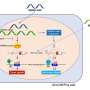
Upon further examination, the study team determined that this particular disorder caused a functional alteration of the AP-2 complex, which is involved in endocytosis, a process by which brain cells recycle small parts of their membrane to create so-called synaptic vesicles that are important in the communication between brain cells. The authors suggest that the key function of the AP2M1 protein is to regulate the balance between excitability and inhibition, a well-established mechanism in epilepsy.
"This is the first time that clinical patient information in a digital format was used to discover a disease gene," said Ethan Goldberg, MD, Ph.D., Assistant Professor of Neurology & Neuroscience and Director of the CHOP ENGIN, a recently launched program that integrates genetic testing into the diagnosis and treatment of children with difficult-to-treat or unexplained epilepsies. "This study emphasizes how the large amount of clinical information that is collected in our patients can facilitate gene discovery and enhance our understanding of a gene's function. Increasingly, knowing how defects in a gene cause seizures allows us to better determine which medications might work and to develop new strategies to treat children with epilepsy. "
More information: Helbig et al, "A recurrent missense variant in AP2M1 impairs clathrin-mediated endocytosis and causes developmental and epileptic encephalopathy." Am J Hum Genet, online May 16, 2019. DOI: 10.1016/j.ajhg.2019.04.001