Potential entry points for Huntington's disease drug discovery

Huntington's disease (HD) is one of the four major neurodegenerative diseases that have been most extensively studied. The clinical symptoms include uncontrolled dancing-mimicking behavior (chorea), as well as cognitive deficiency and psychiatric abnormalities. Since the biochemical activity of the mutant huntingtin protein (mHTT) that causes the disease is uncharacterized, the conventional drug discovery approach, which relied on inhibitors that block the bioactivity of the pathogenic proteins, is not applicable.
Recently, researchers at Fudan University, including Profs. Boxun Lu (School of Life Sciences), Yiyan Fei (School of Informatics Science and Engineering), and Yu Ding (School of Life Sciences), formed a multidisciplinary team to work on this problem. With joint efforts, they have worked out an innovative method of drug discovery: using the autophagosome-tethering compounds (ATTEC) to degrade pathogenic proteins and treat the disease. The team carried out a smartly designed screening featuring a small-molecule microarray and front-edge optical technologies, and managed to identify four small molecule compounds that specifically reduced the protein that causes Huntington's disease.
On October 31, the study was published online under the title "Allele-selective Lowering of Mutant HTT Protein by HTT-LC3 Linker Compounds" in Nature.
"Small molecule glue" helps autophagosomes "engulf" the disease-causing protein
Since the conventional approach is infeasible for mHTT, the team came up with a fundamentally new idea, which was to degrade mHTT by harnessing autophagy, an intracellular protein degradation process. During autophagy, the key protein LC3 is lipidated and expanded to form a double-membrane structure, which then engulfs proteins, lipids, organelles and other degradation cargoes and forms a complete autophagosome. The autophagosomes are then fused with lysosomes and the material engulfed therein is degraded.
However, if enhanced non-specifically, autophagy degrades all proteins engulfed into autophagosomes, including the normal wild-type huntingtin protein that plays a role in neuroprotection. This would be possibly self-defeating.
To identify compounds that only degrade mHTT but not wild-type HTT, the team envisioned a "small molecule glue" functioning as an "autophagosome tethering compound" (ATTEC), which could tether LC3 and mHTT together so that mHTT is engulfed into autophagosomes for degradation. Meanwhile, the ATTEC does not interact with the wild-type HTT protein, leaving it unaffected. Through screening, validation and preliminary search for structurally similar compounds, the team identified four compounds with the desired properties.
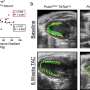
At this point, the identified compounds have the potential of tethering mHTT to autophagosomes without influencing the wild-type HTT, but whether they do function in degrading mHTT as expected needs further validation. The team found that these four compounds significantly reduced mHTT levels in HD mouse neurons, HD patient cells, and HD drosophila models at ~10 to 100 nanomolar concentrations, with little effect on wild-type HTT levels. Excitingly, at least two out of these four compounds are able to cross the blood-brain barrier, and a small dose of intraperitoneal injection would significantly reduce mHTT levels in the cortex and striatum of HD mice, without affecting wild-type HTT levels. They also significantly improved disease-related phenotypes, providing an entry point for the development of oral or injectable drugs for HD.
Such ATTECs were highly challenging to identify. Only one out of about 2000 compounds has the desired properties. Thus, finding it was a major obstacle of this project for a long time. The participation of the Professor Yiyan Fei's group made this possible. Dr. Fei's group developed a new high-throughput compound screening platform based on small molecule microarray (SMM) and oblique-incidence reflectivity difference (OI-RD) technology, which was fast, sensitive, label-free and high-throughput. It could identify the target protein-interacting compounds from a library of thousands of small molecule compounds.
The research team stamped nearly 4000 small-molecule compounds onto a chip and had the target protein flow through the chip. If a sample binds to a specific compound immobilized onto the chip, the molecular layer at the position will thicken, generating a tiny change that can be detected by a sensitive optical method (oblique incident light reflection difference). Using this cutting-edge screening approach, the team found two small molecules that could bind to both LC3 and mHTT proteins, but not to wild-type HTT. After studying a panel of small molecule compounds with similar structures, a total of four ATTECs that bind LC3 and mutant HTT were identified and validated.
Autophagosome tethering compounds may open new windows for drug discovery
The team further explored the intrinsic mechanisms by which these small molecule compounds could distinguish between mutant and wild-type HTT proteins, which were almost identical except in the glutamine repeat (polyQ) length. It turned out that these compounds were bound to excessively long polyQ stretches that only appeared in mHTT.
Based on this, the team realized that the application of these small molecule compounds may reach far beyond the potential treatment of Huntington's disease. Nine human diseases are so called polyQ diseases, because they are caused by specific mutant proteins containing excessively long polyQ. Among them, spinocerebellar ataxia type III (SCA3) is the most common in the Chinese population.
The clinical symptoms include motion discoordination, inability to maintain body posture and balance, accompanied by possible exophthalmos, hyperreflexia, facial muscle twitching, tendon and other symptoms. With SCA3 patient cells provided by Dr. Yimin Sun from Prof. Jian Wang's group at Huashan Hospital affiliated to Fudan University, the team found that these compounds could effectively reduce the level of the mutant ATXN3 protein (with a polyQ length of 74) that causes the disease, without affecting the wild-type ATXN3 (with a polyQ length of 27).
When it came to the future development and applications of this study, Boxun Lu says, "These compounds may not only be effective in the treatment of Huntington's disease, but also applicable to other polyQ diseases. The new concept of drug development using autophagosome-binding compounds (ATTEC) may also be applied to other pathogenic proteins that are undruggable, or even to pathogenic substances that are not proteins, such as organelles or lipids."
More information: Zhaoyang Li et al. Allele-selective lowering of mutant HTT protein by HTT–LC3 linker compounds, Nature (2019). DOI: 10.1038/s41586-019-1722-1