Researchers adapt amyloid technology to catch coronavirus
Two researchers at the University of California, Davis, are adapting their technology for amyloid-based, self-organizing protein scaffolds to combat coronavirus. They hope the technique could be used in diagnostic tests or for virus-neutralizing masks and other protective equipment.
Daniel Cox, professor of physics, and Michael Toney, professor of chemistry, are inventors of a patented technique for using beta amyloid proteins to grow tiny fibrils. The plaques and tangles in the brains of people with Alzheimer's disease and related conditions are described as beta amyloid proteins. The researchers used antifreeze proteins with a similar structure to make their fibrils.
The artificial fibrils act like scaffolds and can be engineered to create new materials with specific functions. Cox and Toney developed the technology for applications such as nanowires and biomaterials. Now their goal is to capture the SARS-CoV-2 virus that causes COVID-19.
"The amyloid polymerization process is responsible for diseases like Alzheimer's, Parkinson's, and mad cow, but can be harnessed in this fashion to make proteins act as controllable molecular building blocks," Cox said.
Platform to help diagnose the new coronavirus
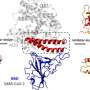
Cox and Toney's prototype centers on protein polymer fibrils engineered to carry a piece of the ACE2 protein. Found on the surface of human cells, ACE2 is the gateway for the SARS-CoV-2 virus to enter cells. They are testing whether the ACE2 fibrils will lock onto the SARS-CoV-2 "spike" protein, which coats the outside of the virus.
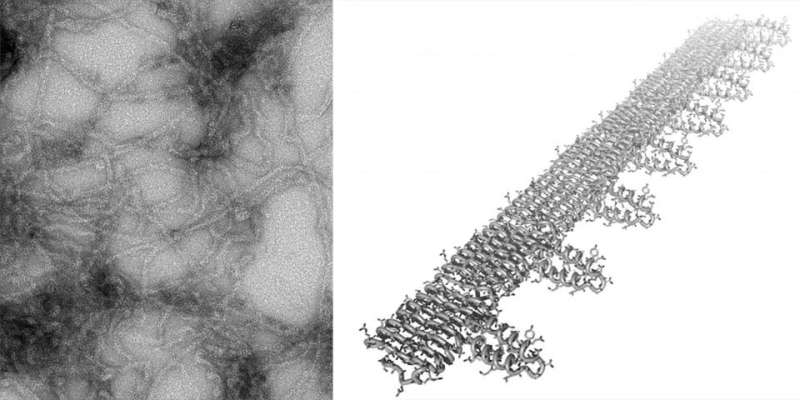
Amyloid proteins naturally self-assemble (or polymerize) into fibrils. To manufacture the ACE2-amyloid proteins, Toney inserted synthetic genes into bacteria. After growing and harvesting the bacteria and isolating the proteins, the purified protein molecules, called monomers, were incubated for several days to encourage fibrils to form. The monomers knit together like Velcro, forming fibrils about 10 micrometers long with the spike-binding piece of ACE2 protein attached along their length.
"Because each monomer has the spike-binding segment of ACE2, these long polymers should serve as really efficient grabbers of the virus," Toney said.
Rapid and sensitive COVID-19 detection
The team hopes their work will contribute to the development of simple, rapid tests to detect the SARS-CoV-2 virus. For example, the polymer fibrils could replace antibodies in existing tests, and boost sensitivity and specificity by capturing more viral proteins and concentrating those proteins for testing, Cox said. The technology could also be used to coat personal protective equipment, such as fabric masks, and boost their effectiveness against the virus, he said.
"We're motivated by the chance to make a difference in protecting people until an effective vaccine is developed," Cox said.
If the prototype polymers succeed, Cox and Toney will need a pharmaceutical partner to manufacture the fibrils and test them in clinical trials. And before any test is deployed, it must be approved by the Food and Drug Administration.