New drug targets for lethal brain cancer discovered
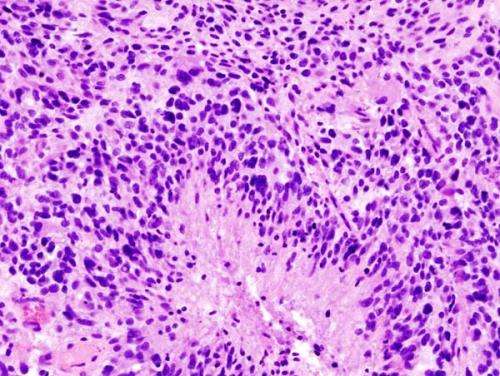
More than 200 genes with novel and known roles in glioblastoma—the most aggressive type of brain cancer—offer promising new drug targets. Researchers from the Wellcome Sanger Institute, Addenbrooke's Hospital and their collaborators engineered a new mouse model to show for the first time how a mutation in the well-known cancer gene, EGFR initiates glioblastoma, and works with a selection from more than 200 other genes to drive the cancer.
The results, published today in Genome Biology present the first mouse model of its kind, which is available for the research community to advance new treatments for this lethal form of brain cancer.
Glioblastoma is an aggressive form of brain cancer. It is treated with surgery followed by chemotherapy or radiotherapy, however glioblastoma cells can evade treatment and tumors return. The prognosis is poor—the average patient survives for 12-18 months following diagnosis.
New, targeted treatments and immunotherapies are currently being developed to help glioblastoma patients. It is still not known exactly why glioblastomas begin to grow.
In a new study, researchers from the Wellcome Sanger Institute and their collaborators created a new mouse model with glioblastoma to investigate which genes were implicated in the cancer.
The model showed that the well-known cancer gene, EGFR (epidermal growth factor receptor) can alone initiate the brain tumors to grow in mice, resulting in tumors that were highly representative of human glioblastomas.
Dr. Imran Noorani, a corresponding author previously from the Wellcome Sanger Institute, and now based at Addenbrooke's Hospital and the University of Cambridge, said: "We have created a new mouse model for studying the lethal human brain cancer, glioblastoma. For the first time, we showed that the familiar cancer gene, EGFR is capable of initiating glioblastoma and we identified new driver genes, whose potential for therapeutic targeting deserves further exploration."
To identify which genes help EGFR to drive the cancer, the team used the PiggyBac transposon technique—a small section of DNA inserted into different parts of the genome to introduce mutations. This revealed more than 200 known and novel mutations in tumor suppressor genes that were working with EGFR to drive brain tumor growth, many of which present new drug targets.
The team compared the results with human genome sequences from glioblastoma patients and uncovered many genetic mutations found in both humans and mice. Human genomic data contains many mutations implicated in glioblastoma, without clear indication of which specific mutations drive the cancer. With the new mouse model, the team were able to narrow down on which mutations drive glioblastoma, which will focus future drug development.
Professor Allan Bradley, previously Director of the Wellcome Sanger Institute, and now Chief Scientific Officer of Kymab and Professor in the Department of Medicine, University of Cambridge, said: "Glioblastoma patients urgently require new, targeted therapies. Unfortunately, glioblastoma tumors can become highly resistant to therapies that target specific molecules, as there are many other genetic drivers that can 'take over' progressing the cancer. This new mouse model provides the missing link to translate findings from new potential treatments tested on mice to clinical trials."
More information: Imran Noorani et al. (2020) PiggyBac mutagenesis and exome sequencing identify genetic driver landscapes and potential therapeutic targets of EGFR-mutant gliomas. Genome Biology. DOI: 10.1186/s13059-020-02092-2