Genomic markers of midostaurin drug sensitivity in leukemia patients
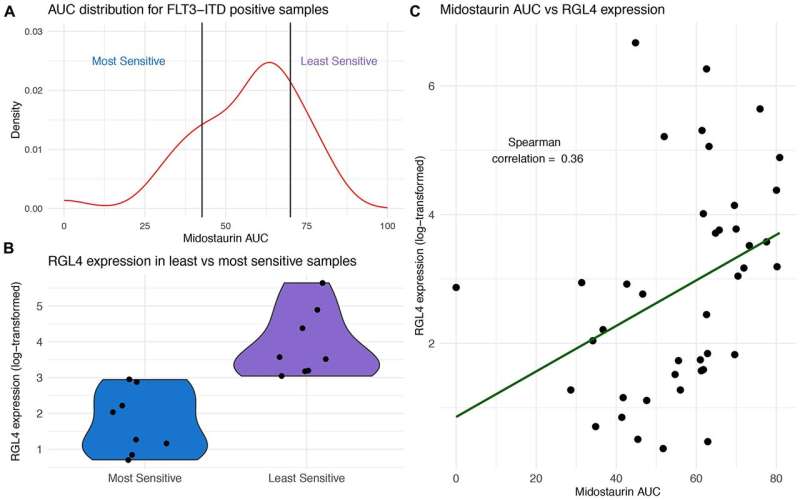
Researchers have identified acute myeloid leukemia as a heterogeneous malignancy with the most common genomic alterations in NPM1, DNMT3A and FLT3. Midostaurin was the first FLT3 inhibitor FDA approved for AML and is standard of care for FLT3 mutant patients undergoing induction chemotherapy. As there is a spectrum of response, the researchers hypothesized that biological factors beyond FLT3 could play a role in drug sensitivity and that select FLT3-ITD negative samples may also demonstrate sensitivity.
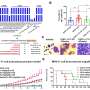
The authors of the study, published in Oncotarget, performed an ex vivo drug sensitivity screen on primary and relapsed AML samples with corresponding targeted sequencing and RNA sequencing. They observed a correlation between FLT3-ITD mutations and midostaurin sensitivity as expected and observed KRAS and TP53 mutations correlating with midostaurin resistance in FLT3-ITD negative samples. Further, they identified genes differentially expressed in sensitive vs. resistant samples independent of FLT3-ITD status.
Within FLT3-ITD mutant samples, over-expression of RGL4, oncogene and regulator of the Ras-Raf-MEK-ERK cascade, distinguished resistant from sensitive samples.
Dr. Mara W. Rosenberg from The Oregon Health and Science University said, "Acute myeloid leukemia (AML) is a heterogeneous malignancy, most commonly affecting individuals ≥60 years of age." FLT3 mutations occur in approximately 30% of de novo AML cases, of which 25% are ITD mutations and 5% are tyrosine kinase domain point mutations. Per the 2017 European Leukemia Network Guidelines, a complete diagnostic work-up should include screening for the presence of FLT3 mutations, as well as mutant-to-wild-type allelic ratios.
There are many past and present clinical trials examining the activity of tyrosine kinase inhibitors against FLT3 mutant AML, including sunitinib, midostaurin, lestaurtinib, sorafenib, ponatinib, crenolanib, gilteritinib, and quizartinib. Factors predictive of the development of resistance include the initial presence of multiple leukemic clones, low FLT3-mutant allelic ratio, or additional primary mutations in the FLT3 kinase domain.
The authors hypothesized that there are additional genomic alterations and gene expression changes outside of FLT3-ITD mutations that can influence AML sample resistance or sensitivity to midostaurin and aimed to further characterize these factors.
The Rosenberg Research Team writes, "We identify genomic alterations that correlate with midsotaurin response independent of FLT3-ITD status, propose that Ras-Raf-MEK-ERK inhibition in combination therapy could limit resistance to midostaurin, and suggest that within the overall AML population there may be therapeutic benefit of midostaurin in patients with certain expression profiles."
More information: Mara W. Rosenberg et al, Genomic markers of midostaurin drug sensitivity in FLT3 mutated and FLT3 wild-type acute myeloid leukemia patients, Oncotarget (2020). DOI: 10.18632/oncotarget.27656