Computer model makes strides in search for COVID-19 treatments
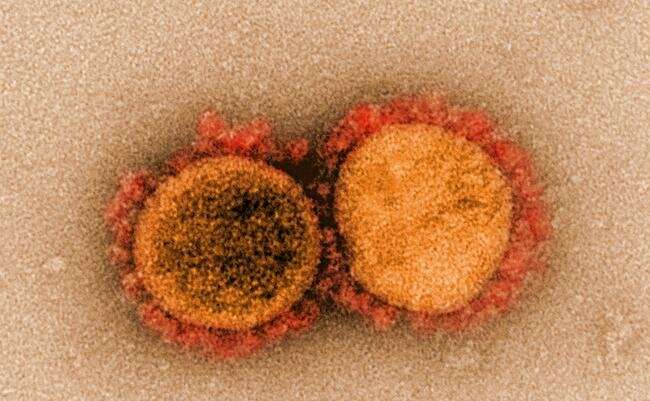
A new deep-learning model that can predict how human genes and medicines will interact has identified at least 10 compounds that may hold promise as treatments for COVID-19.
All but two of the drugs are still considered investigational and are being tested for effectiveness against hepatitis C, fungal disease, cancer and heart disease. The list also includes the approved drugs cyclosporine, an immunosuppressant that prevents transplant organ rejection, and anidulafungin, an antifungal agent.
The discovery was made by computer scientists, meaning much more work needs to be done before any of these medications would be confirmed as safe and effective treatments for people infected with SARS-CoV-2. But by using artificial intelligence to arrive at these options, the scientists have saved pharmaceutical and clinical researchers the time and money it would take to search for potential COVID-19 drugs on a piecemeal basis.
"When no one has any information on a new disease, this model shows how artificial intelligence can help solve the problem of how to consider a potential treatment," said senior author Ping Zhang, assistant professor of computer science and engineering and biomedical informatics at The Ohio State University.
The researchers noted in the paper that a few of the repurposing candidates the model generated have already been studied for their potential use in COVID-19 patients.
"Great minds think alike—some lead compounds identified by machine intelligence coincide with later discoveries by human intelligence," Zhang said.
The research is published today (Feb. 1) in Nature Machine Intelligence.
Zhang and colleagues had completed the model's design in May 2020, just as the first papers detailing how COVID-19 patients' genes responded to the virus were published. The new information provided an important test for the computer model, which the researchers call "DeepCE"—pronounced "Deep Sea."
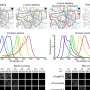
To make predictions about how genes and medicines will interact and yield drug repurposing candidates, DeepCE relies on two primary sources of publicly available data: L1000, a National Institutes of Health-funded repository of human cell-line data showing how gene expression changes in response to drugs, and DrugBank, which contains information on the chemical structures and other details on about 11,000 approved and investigational drugs.
L1000 displays side-by-side cell-line comparisons of standard gene expression activity with gene expression changes produced by interactions with specific drugs. The cell lines represent diseases, such as melanoma, and organs, like kidneys and lungs. It is an ongoing project, with data being added as experiments in animals or humans supplement the gene expression profiles produced in cell-line experiments.
The Ohio State researchers trained the DeepCE model by running all of the L1000 data through an algorithm against specific chemical compounds and their dosages. To fill in data gaps, the model converts chemical compound descriptions into figures, allowing for automatic consideration of their separate components' effects on genes. And for genes not represented in L1000, the team used a deep learning approach called an "attention mechanism" to increase the model's "learned" sample of gene-chemical compound interactions, which improves the framework's performance.
"This way, the output demonstrates multitask learning—we can predict gene expression values for new chemicals not from one cell to one cell, but automatically predict the role of a drug on different cell lines and different genes," said Zhang, who leads the Artificial Intelligence in Medicine Lab and is a core faculty member in the Translational Data Analytics Institute at Ohio State. "We can use the computer to simulate drug-induced gene expression. This provides real value."
"The story should stop here—this is where we were during spring break. But then COVID-19 arrived, and we hoped our research could help, so we did a special case study for COVID-19 drug repurposing."
The team applied DeepCE's gene expression prediction matrix—focusing on data from lung and airway cell lines and the entire DrugBank catalog of compounds—to the genetic information provided from the early COVID-19 papers and additional government data. The COVID-19 data demonstrated how human gene expression had responded to being infected with SARS-CoV-2, creating a "disease signature."
"Based on the known gene expression changes that have occurred and been identified with known drugs, we apply that to the gene expression in question—in this case, compounds that are being studied but are not yet experimented in L1000. We put such predicted 'drug signatures' against the COVID-19 patient profiles on a population level," Zhang said.
"Once you can identify both signatures, the job is easy. Wherever we find the disease and a drug show opposite gene expression profiles, suggesting the drug would reverse the effects of the disease, you have found a drug that may treat the disease."
This model complements a drug-repurposing model Zhang described in a recent paper that simulates clinical trials using observational clinical data.
"I want to put together a research agenda using all the different data resources for drug repurposing and drug-disease associations from multiple perspectives and connect with researchers who can collaborate with us to find new drugs for diseases—including unknown diseases," Zhang said.
More information: Pham, TH., Qiu, Y., Zeng, J. et al. A deep learning framework for high-throughput mechanism-driven phenotype compound screening and its application to COVID-19 drug repurposing. Nat Mach Intell (2021). doi.org/10.1038/s42256-020-00285-9