Reading minds with ultrasound: A less-invasive technique to decode the brain's intentions
What is happening in your brain as you are scrolling through this page? In other words, which areas of your brain are active, which neurons are talking to which others, and what signals are they sending to your muscles?
Mapping neural activity to corresponding behaviors is a major goal for neuroscientists developing brain-machine interfaces (BMIs): devices that read and interpret brain activity and transmit instructions to a computer or machine. Though this may seem like science fiction, existing BMIs can, for example, connect a paralyzed person with a robotic arm; the device interprets the person's neural activity and intentions and moves the robotic arm correspondingly.
A major limitation for the development of BMIs is that the devices require invasive brain surgery to read out neural activity. But now, a collaboration at Caltech has developed a new type of minimally invasive BMI to read out brain activity corresponding to the planning of movement. Using functional ultrasound (fUS) technology, it can accurately map brain activity from precise regions deep within the brain at a resolution of 100 micrometers (the size of a single neuron is approximately 10 micrometers).
The new fUS technology is a major step in creating less invasive, yet still highly capable, BMIs.
"Invasive forms of brain-machine interfaces can already give movement back to those who have lost it due to neurological injury or disease," says Sumner Norman, postdoctoral fellow in the Andersen lab and co-first author on the new study. "Unfortunately, only a select few with the most severe paralysis are eligible and willing to have electrodes implanted into their brain. Functional ultrasound is an incredibly exciting new method to record detailed brain activity without damaging brain tissue. We pushed the limits of ultrasound neuroimaging and were thrilled that it could predict movement. What's most exciting is that fUS is a young technique with huge potential—this is just our first step in bringing high performance, less invasive BMI to more people."
The new study is a collaboration between the laboratories of Richard Andersen, James G. Boswell Professor of Neuroscience and Leadership Chair and director of the Tianqiao and Chrissy Chen Brain-Machine Interface Center in the Tianqiao and Chrissy Chen Institute for Neuroscience at Caltech; and of Mikhail Shapiro, professor of chemical engineering and Heritage Medical Research Institute Investigator. Shapiro is an affiliated faculty member with the Chen Institute.
A paper describing the work appears in the journal Neuron on March 22.
In general, all tools for measuring brain activity have drawbacks. Implanted electrodes (electrophysiology) can very precisely measure activity on the level of single neurons, but, of course, require the implantation of those electrodes into the brain. Non-invasive techniques like functional magnetic resonance imaging (fMRI) can image the entire brain but require bulky and expensive machinery. Electroencephalography (EEGs) does not require surgery but can only measure activity at low spatial resolution.
Ultrasound works by emitting pulses of high frequency sound and measuring how those sound vibrations echo throughout a substance, such as various tissues of the human body. Sound travels at different speeds through these tissue types and reflects at the boundaries between them. This technique is commonly used to take images of a fetus in utero, and for other diagnostic imaging.
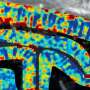
Ultrasound can also "hear" the internal motion of organs. For example, red blood cells, like a passing ambulance, will increase in pitch as they approach the source of the ultrasound waves, and decrease as they flow away. Measuring this phenomenon allowed the researchers to record tiny changes in the brain's blood flow down to 100 micrometers (on the scale of the width of a human hair).
"When a part of the brain becomes more active, there's an increase in blood flow to the area. A key question in this work was: If we have a technique like functional ultrasound that gives us high-resolution images of the brain's blood flow dynamics in space and over time, is there enough information from that imaging to decode something useful about behavior?" Shapiro says. "The answer is yes. This technique produced detailed images of the dynamics of neural signals in our target region that could not be seen with other non-invasive techniques like fMRI. We produced a level of detail approaching electrophysiology, but with a far less invasive procedure."
The collaboration began when Shapiro invited Mickael Tanter, a pioneer in functional ultrasound and director of Physics for Medicine Paris (ESPCI Paris Sciences et Lettres University, Inserm, CNRS), to give a seminar at Caltech in 2015. Vasileios Christopoulos, a former Andersen lab postdoctoral scholar (now an assistant professor at UC Riverside), attended the talk and proposed a collaboration. Shapiro, Andersen, and Tanter then received an NIH BRAIN Initiative grant to pursue the research. The work at Caltech was led by Norman, former Shapiro lab postdoctoral fellow David Maresca (now assistant professor at Delft University of Technology), and Christopoulos. Along with Norman, Maresca and Christopoulos are co-first authors on the new study.
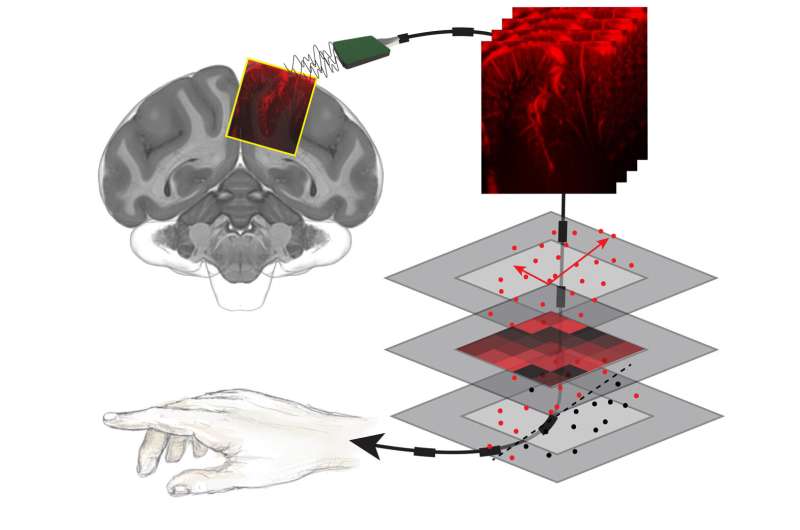
The technology was developed with the aid of non-human primates, who were taught to do simple tasks that involved moving their eyes or arms in certain directions when presented with certain cues. As the primates completed the tasks, the fUS measured brain activity in the posterior parietal cortex (PPC), a region of the brain involved in planning movement. The Andersen lab has studied the PPC for decades and has previously created maps of brain activity in the region using electrophysiology. To validate the accuracy of fUS, the researchers compared brain imaging activity from fUS to previously obtained detailed electrophysiology data.
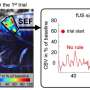
Next, through the support of the T&C Chen Brain-Machine Interface Center at Caltech, the team aimed to see if the activity-dependent changes in the fUS images could be used to decode the intentions of the non-human primate, even before it initiated a movement. The ultrasound imaging data and the corresponding tasks were then processed by a machine-learning algorithm, which learned what patterns of brain activity correlated with which tasks. Once the algorithm was trained, it was presented with ultrasound data collected in real time from the non-human primates.
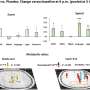
The algorithm predicted, within a few seconds, what behavior the non-human primate was going to carry out (eye movement or reach), direction of the movement (left or right), and when they planned to make the movement.
"The first milestone was to show that ultrasound could capture brain signals related to the thought of planning a physical movement," says Maresca, who has expertise in ultrasound imaging. "Functional ultrasound imaging manages to record these signals with 10 times more sensitivity and better resolution than functional MRI. This finding is at the core of the success of brain-machine interfacing based on functional ultrasound."
"Current high-resolution brain-machine interfaces use electrode arrays that require brain surgery, which includes opening the dura, the strong fibrous membrane between the skull and the brain, and implanting the electrodes directly into the brain. But ultrasound signals can pass through the dura and brain non-invasively. Only a small, ultrasound-transparent window needs to be implanted in the skull; this surgery is significantly less invasive than that required for implanting electrodes," says Andersen.
Though this research was carried out in non-human primates, a collaboration is in the works with Dr. Charles Liu, a neurosurgeon at USC, to study the technology with human volunteers who, because of traumatic brain injuries, have had a piece of skull removed. Because ultrasound waves can pass unaffected through these "acoustic windows," it will be possible to study how well functional ultrasound can measure and decode brain activity in these individuals.
More information: Neuron (2021). DOI: 10.1016/j.neuron.2021.03.003