Researchers identify the proteins that cause intestinal disease
Researchers from Tel Aviv University have created an artificial intelligence platform that can identify the specific proteins that allow bacteria to infect the intestines—a method that paves the way for the creation of smart drugs that will neutralize the proteins and prevent disease, without the use of antibiotics. Participating in the study, which was published in the prestigious journal Science, were Ph.D. student Naama Wagner and Prof. Tal Pupko, head of the Shmunis School of Biomedicine and Cancer Research at the Faculty of Life Sciences and the new Center for Artificial Intelligence & Data Science at Tel Aviv University. The international partners in the study included researchers from Imperial College (led by Prof. Gad Frankel) and the Institute for Cancer Research in London, as well as from the Technical University and the National Center for Biotechnology in Madrid.
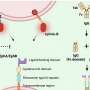
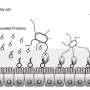
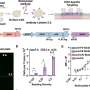
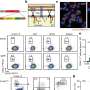
Intestinal diseases are caused by pathogenic bacteria that attach to our intestinal cells. Once attached, the bacteria use a kind of molecular syringe to inject intestinal cells with proteins called "effectors." These effectors work together to take over healthy cells, like hackers that take over computer servers using a combination of lines of code. However, until now scientists have not known what protein combination it is that cracks the cell's defense mechanisms. Now, the Tel Aviv University researchers' artificial intelligence platform has identified novel effectors in the bacteria, which have been experimentally tested and validated. Subsequently, laboratory experiments conducted in London successfully predicted the protein combinations that lead to the pathogenic bacteria taking over the intestines.
"In this study, we focused on a bacterium that causes intestinal disease in mice, a relative of the E. coli bacteria that cause intestinal disease in humans, so as not to work directly with the human pathogen," explains Ph.D. student Naama Wagner. "The artificial intelligence we created knows how to predict effectors in a variety of pathogenic bacteria, including bacteria that attack plants of economic importance. Our calculations were made possible by advanced machine-learning tools that use the genomic information of a large number of bacteria. Our partners in England proved experimentally that the learning was extremely accurate and that the effectors we identified are indeed the weapons used by the bacteria."
"Pathogenic bacteria are treated with antibiotics," says Prof. Tal Pupko. "But antibiotics kill a large number of species of bacteria, in the hope that the pathogenic bacteria will also be destroyed. So antibiotics are not a rifle but a cannon. Moreover, the overuse of antibiotics leads to the development of antibiotic-resistant bacteria, a worldwide problem that is getting worse. Understanding the molecular foundation of the disease is a necessary step in the development of drugs that are smarter than antibiotics, which will not harm the bacterial population in the intestines at all. This time we discovered the effectors of gut bacteria that attack rodents, but this is just the beginning. We are already working on detecting effectors in other bacteria in an attempt to better understand how they carry out their mission in the target cells they are attacking."
More information: David Ruano-Gallego et al, Type III secretion system effectors form robust and flexible intracellular virulence networks, Science (2021). DOI: 10.1126/science.abc9531