New insights into the virulence arsenal of a common hospital pathogen
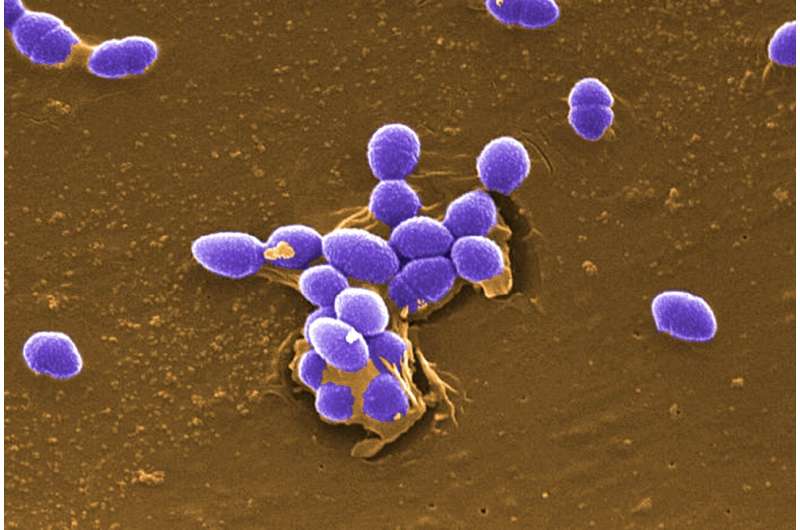
Each pathogen relies on a unique set of strategies and surface & secreted molecules that enable it to colonize and infect host cells, which are collectively known as "virulence factors."
Studying these strategies helps scientists unravel the disease mechanism, identify diagnostic markers and expose new therapeutic targets.
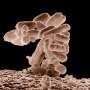
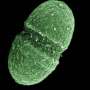
Recently, researchers from Children's Cancer Hospital 57357 and The British University in Egypt teamed up on a study revealing new insights on virulence factors of Enterococcus faecalis, a common hospital-acquired pathogen.
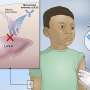
Under normal conditions, E. faecalis microbes live harmlessly inside our gastrointestinal tract, and might even have healthy effects on the GI system. However, when the immune system is compromised or when the bacteria are in an organ to which they do not usually have access, they seize the opportunity by deploying their virulence molecules causing several diseases, especially urinary tract infections. These infections usually sweep in hospitals, complicating the treatment of patients with serious conditions, like cancer.
In this study, the team collected 60 urinary samples of E. faecalis from Egyptian clinical laboratories after obtaining the required ethical approvals.
Thereafter, they cultured the bacteria in a proper environment in the lab that allows it to grow and spread out its virulence arsenal, so the team could analyze it. Concurrently, the team extracted the genetic material from the samples, looking for the concorded genes of these virulence features.
"Correlating the virulence factors to its genetic basis is essential to help us zoom into the main diagnostic genes, and might also reveal therapeutic targets." Says Ramy Aziz head of immunology and microbiology research program in CCHE 57, and the corresponding author on the paper.
The results confirm that the ability of E. faecalis to adhere to the host tissues is an essential strategy for its pathogenesis. All isolates were able to form a thin, sticky layer of bacteria known as a biofilm, which can irreversibly attach to the inner wall of the urinary tract, enabling bacteria to resist flushing by urine. Moreover, all isolates harbored two adherence-related genes.
Releasing cytolysin toxin that can harm and rupture red blood cells is another virulent feature of E. faecalis that can increase the infection severity. The lysis of blood cells known as hemolysis was significantly correlated with the presence of the cytolysin-encoding gene (cyl), which suggests using cyl as a biomarker, the researchers conclude.
However, not all virulence features could be easily inferred through the genes. For instance, the activity of the bacterial gelatinase enzyme that plays an important role in spreading the virus through the body was absent in most isolates, although the gelatinase gene (gelE) was present in almost all of them.
"Some genes might be present but are not active, and [some virulence features are controlled by a complex network of genes," Aziz said.
For future research, Aziz and his team intend to sequence the whole genome of the E. faecalis isolates with the hope of uncovering new valid biomarkers that could aid in accurate timely diagnosis.
The study was published in the May issue of Infection and Drug Resistance.
More information: Yomna A Hashem et al, Phenotype–Genotype Correlations and Distribution of Key Virulence Factors in Enterococcus faecalis Isolated from Patients with Urinary Tract Infections, Infection and Drug Resistance (2021). DOI: 10.2147/IDR.S305167