Classifying EGFR mutations by structure offers better way to match non-small cell lung cancer patients to treatments
Researchers from The University of Texas MD Anderson Cancer Center have discovered that grouping epidermal growth factor receptor (EGFR) mutations by structure and function provides an accurate framework to match patients with non-small cell lung cancer (NSCLC) to the right drugs. The findings, published today in Nature, identify four subgroups of mutations and introduce a new strategy for testing tyrosine kinase inhibitors (TKIs), as well as instant clinical opportunities for approved targeted therapies.
"More than 70 different EGFR mutations have been identified in patients, but drugs have only been approved for a handful of them. One of the immediate implications of our research is the discovery that therapies we already have may work for many of these mutations. For some mutations, older drugs may actually work better, and for other mutations, newer drugs work better," said John Heymach, M.D., Ph.D., chair of Thoracic/Head & Neck Medical Oncology and senior author of the study. "Right now, in the absence of guidance, clinicians often use the newest treatment for all EGFR mutations. This model can help us pick better therapies for patients immediately and hopefully develop better drugs for specific subgroups of mutations."
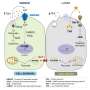
First-, second- and third-generation TKIs use different mechanisms to target the EGFR protein. Heymach and his team found that drugs work better for certain subgroups based on how the mutations within a given group functionally impact the drug-binding pocket on the protein.
The four EGFR-mutant NSCLC subgroups identified by the team are:
- Classical-like mutations, with little to no impact on drug binding
- T790M-like mutations, which contain at least one mutation in the hydrophobic cleft and often are acquired after resistance to a first-generation targeted therapy
- Exon 20 loop insertion mutations, characterized by insertions of additional amino acids in the loop after the C-terminal end of the αC-helix in exon 20
- P-loop αC-helix compression (PACC) mutations on the interior surface of the ATP binding pocket or C-terminal end of the αC-helix
The current approach to testing new drugs in EGFR-mutant NSCLC is based on exon number, which indicates where the mutation occurs within a linear portion of the DNA. Grouping mutations by exon has produced mostly heterogeneous results in clinical studies and laboratory models, which the authors note seems to indicate a poor correlation between exon number and drug sensitivity or resistance.
"Within a given exon, mutations vary widely. We organized mutations based on how they impact the EGFR structure and drug binding instead, which allows for testing a drug across a whole group of mutations that are structurally similar at the same time," Heymach said. "We believe this could become the new standard approach for classifying and describing mutations and then pairing them with the right drug."
Big data reveals diversity in atypical mutations
Mutations in the EGFR protein are present in about 15% of NSCLCs in North America and about 30 to 40% in Asia. Overall, more than 70 different types of EGFR mutations exist. "Classical" mutations tend to respond well to FDA-approved targeted therapies, but effective therapies and guidelines for the remaining "atypical" mutations have been lacking.
For this study, the researchers analyzed data from 16,175 patients with EGFR-mutant NSCLC from five different patient databases. Primary and co-occurring mutations were recorded for 11,619 patients. Of those, 67.1% had classical EGFR mutations, 30.8% had atypical EGFR mutations and 2.2% had both.
One of the key databases to provide detailed molecular and outcome information for the study was the Genomic Marker-Guided Therapy Initiative (GEMINI), a big data project of the Lung Cancer Moon Shot, part of MD Anderson's Moon Shots Program, a collaborative effort designed to accelerate the development of scientific discoveries into clinical advances that save patients' lives.
For both classical and atypical EGFR mutations, the team analyzed the time to treatment failure (TTF), an indication of how quickly a cancer becomes resistant to therapy. The researchers found a shorter TTF and lower overall survival for patients with atypical mutations regardless of treatment type. Patients with classical mutations treated with first- and third-generation TKIs had a longer TTF.
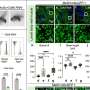
The researchers then created a panel of 76 cell lines with EGFR mutations and screened those cell lines against 18 EGFR inhibitors, which revealed the four distinct subgroups. Comparing the correlation to drug sensitivity by subgroup, versus exons, showed that the structure-based subgroups were more predictive than exon-based groups. The subgroup approach was further validated by machine learning to analyze data by classification and regression trees.
Classical-like mutations were sensitive to all classes of TKIs, particularly third-generation TKIs. Exon 20 loop insertion mutations remained the most heterogeneous subgroup, with certain mutations responding best to second-generation TKIs. T790M-like mutations were sensitive to ALK and PKC inhibitors, with some mutations retaining sensitivity to third-generation TKIs. PACC mutations were most sensitive to second-generation TKIs.
"Proteins aren't linear, so grouping mutations by exon didn't seem an intuitive approach to me when I started thinking about how to match the right drug to the right mutation seen in patients," said Jacqulyne Robichaux, Ph.D., assistant professor of Thoracic/Head & Neck Medical Oncology Research and lead author of the study. "Proteins are three dimensional, and this led us to investigate if there were areas of the proteins that correlate with drug sensitivity when mutated, which is what we found. These subgroups share properties in their structure that directly relate to their function and retrospectively predicted patient outcomes better than the traditional approach."
Further emphasis on the role of next-generation sequencing and future studies
The study also highlights the importance of biomarker testing for all patients with a new diagnosis or recurrence of NSCLC. Current next-generation sequencing methods have the ability to detect the full spectrum of known oncogenic driver EGFR mutations, virtually all of which fall into one of the four structure-based subgroups. The authors note that this is especially important for rare mutations, which are more difficult to study through a traditional clinical trial approach based on individual mutations. Future prospective studies will help refine and inform the subgroup framework.
"This is an important advance for patients because, right now, there is no FDA-approved targeted therapy for the majority of EGFR mutations, leaving clinicians in the dark as to what drug to use for which mutation," Heymach said. "Now, based on the structural group in which the mutation falls, we can better match the best drug for a given mutation. Going forward, this may also help focus drug development efforts, by testing drugs against an entire group of mutations that are structurally similar, rather than against individual mutations."
More information: Structure-based classification predicts drug response in EGFR-mutant NSCLC, Nature (2021). DOI: 10.1038/s41586-021-03898-1 , www.nature.com/articles/s41586-021-03898-1