More accurately identifying colorectal cancer through microbiome composition
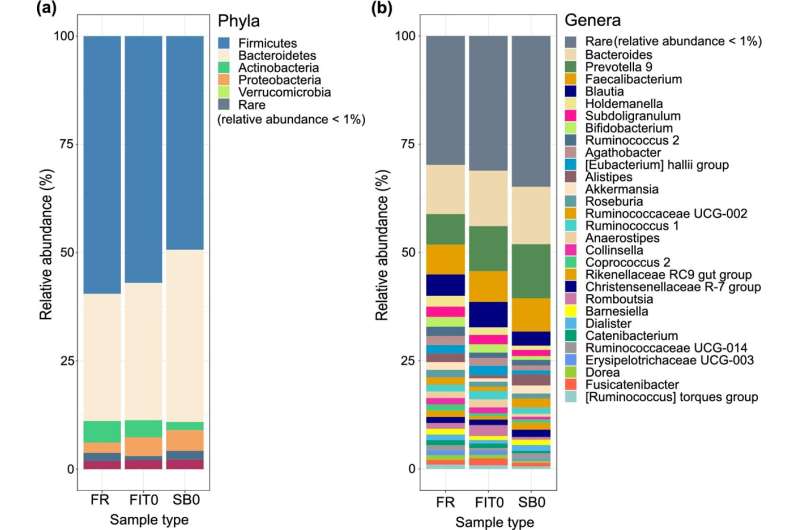
In order to study colorectal cancer more efficiently, microbiome scientists from the University of Tartu have been able to characterize microbiome composition from sample tubes used in colorectal cancer screening for testing fecal occult blood.
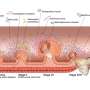
Epidemiological data suggest that the incidence of colorectal cancer is expected to increase 60% by 2030 due to population aging and the growing popularity of Western diets and lifestyles. Many countries have started inviting individuals to participate in population-based screening programs to increase early cancer detection by analyzing fecal blood via a fecal immunochemical test (FIT), followed by colonoscopy.
"Current screening programs, however, face multiple challenges, including low participation rates, low sensitivity for pre-cancerous or early-stage cancer, and false positive and false-negative results among others. Consequently, new highly specific, inexpensive, and sensitive noninvasive screening tests that improve the detection of precancerous colorectal lesions and cancer are urgently needed to reduce the incidence and mortality of the disease," said Kertu Liis Krigul, the first author of the paper.
Accumulating evidence indicates that the microbiome substantially contributes to the development of colorectal cancer. Therefore, complementing fecal occult blood tests with gut microbiota analysis could improve the detection of colorectal cancer.
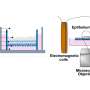
In this study, the researchers tested FIT tubes used in national colorectal cancer screening programs on the volunteers and found that these are also suitable for microbiome analysis. However, the actual ability to detect cancer-specific bacterial signatures with this collection method will now be studied in patients with and without CRC.
"The detection of colorectal cancer with additional microbiome-based biomarkers could potentially make colorectal cancer diagnostics more sensitive and cost-efficient," said Krigul. Fecal samples from colorectal cancer screening programs could be an excellent resource for biomarker discovery, which might lead to earlier detection of cancer or prior to its onset (in a precancerous state).
Authors assume that the stool samples from FIT tubes could also be used as a resource for biomarker discovery and for studying the role of the gut microbiome in other digestive diseases previously associated with the gut microbiome, such as pancreatic cancer, inflammatory bowel disease and cholangitis, as well as in population studies, in which samples are often sent via post and using fresh-frozen samples is not possible.
"Obtaining accurate microbiome profiles from FIT tubes used in multiple European screening programs, therefore, was a crucial step for new study possibilities and future biomarker discovery easing the collection of the microbiome," said Krigul.
The study is published in Scientific Reports.
More information: Kertu Liis Krigul et al, Using fecal immunochemical tubes for the analysis of the gut microbiome has the potential to improve colorectal cancer screening, Scientific Reports (2021). DOI: 10.1038/s41598-021-99046-w