More effective COVID-19 treatment with immune modulatory drugs
A recent study published in the Proceedings of the National Academy of Sciences by a team of researchers at the University of Colorado Anschutz Medical Campus significantly advances the understanding of a key aspect of the immune system during COVID-19: the interferon response.
Interferons (IFNs) are signaling proteins produced by a host cell to activate the antiviral defenses within the body. If the immune response, including the production of IFNs, is unable to clear the virus, the immune system continues to fight. However, a prolonged and exacerbated immune response can cause organ damage and even death.
The collaborative team led by Joaquin Espinosa, Ph.D., executive director of the Linda Crnic Institute for Down Syndrome and professor at the University of Colorado School of Medicine used blood samples and data from patients hospitalized with COVID-19 through a partnership with UCHealth and Children's Hospital Colorado. The samples were provided through the CU COVID Biobank led by Thomas Flaig, MD, vice chancellor of research at the CU Anschutz Medical Campus, and subjected to a deep analysis using the latest 'omics technologies as part of a multidisciplinary effort led by Espinosa and Flaig known as the COVIDome Project. Omics technologies analyze the genome, transcriptome, proteome, and metabolome of biological samples. Importantly, this large dataset is made openly available through the COVIDome Explorer portal, a user-friendly online tool that has been employed by 1500+ users across 60+ countries since its launch in November 2020.
The development of the COVIDome dataset was modeled after the Crnic Institute Human Trisome Project (HTP) which was launched in 2016. The HTP is co-organized and funded by the Global Down Syndrome Foundation and is one of the world's largest studies aimed at understanding co-occurring conditions of Down syndrome. Notably, individuals with Down syndrome are at very high risk of developing severe COVID-19.
This research is the first of its kind in using a deep multi-omics approach to investigate the action of 12 different IFNs in COVID-19 patients. These latest findings built on the team's previous published work and provide a new model for staging the course of COVID-19 pathology.
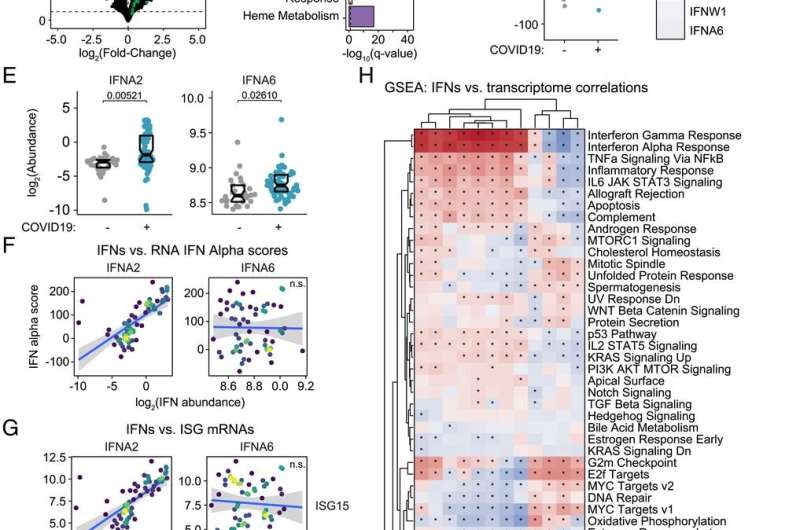
"We were able to measure 12 different IFNs and track their action throughout the course of COVID-19 by defining associations with thousands of RNAs, proteins, antibodies, metabolites, and immune cells measured in the same blood samples. There are no other datasets that have compared that many different measurements to 12 different IFNs in this way," explains the lead author of the paper, Matthew Galbraith, Ph.D., associate research professor at the University of Colorado School of Medicine and head of the Data Sciences Program at the Crnic Institute,
According to co-author Kelly Sullivan, Ph.D., assistant professor at the University of Colorado School of Medicine, there are clear implications in the results that resolve ongoing controversies in the field of COVID-19 research. "There is a big debate about the protective versus harmful effects of IFNs during the immune response in COVID-19. Part of the controversy is driven by the assumption that all IFNs work fairly similarly, but we found massive specialization in IFN action, with different IFNs being associated with different pathological processes in COVID-19 disease."
"These findings have important therapeutic implications, because both pro and anti-IFN therapies have been tested in COVID-19, with mixed results. The key message from this work is that it is more complicated, as not all IFNs are created equal, and that different IFNs likely modulate diverse aspects of the immune response at distinct stages of COVID-19. Therefore, immune modulatory strategies must consider the stage of disease and the IFN subtypes being produced by the patient," explains Tell Bennett, MD, coauthor of the study and director of the Informatics Core of the Colorado Clinical and Translational Sciences Institute involved in data generation and analysis.
The FDA has approved the emergency use authorization for the drug baricitinib for patients hospitalized with severe COVID-19. Baricitinib is a janus kinase inhibitor (JAK inhibitor) that suppresses the action of IFNs and other inflammatory processes. However, it is not clear which patients would benefit the most from this therapy and why suppressing the immune response in this way is often beneficial. Dr. Espinosa believes the results from this study are a significant step toward better understanding how to treat severe COVID-19 with JAK inhibitors and other immune modulatory medicines.
Co-author Elena Hsieh, MD, who specializes in pediatric allergy and immunology at Children's Hospital Colorado and is also a member of the international COVID Human Genetic Effort, sees potential benefits of these findings even beyond therapeutic interventions. "I think this research brings us closer to the identification of a predictive marker about who is more at risk for severe COVID-19 and who isn't. If physicians could do a 'multi-IFN test' to measure the specific IFN signatures along the course of COVID-19, and determine which patients at what stage of their disease would benefit from pro or anti-IFN therapy, it will support a precision medicine approach to COVID-19 management."
While follow up mechanistic studies will be needed to fully define the impacts of specialized IFN action during COVID-19, this work provides a solid foundation for future research toward better prevention and treatment of infectious diseases overall.
More information: Matthew D. Galbraith et al, Specialized interferon action in COVID-19, Proceedings of the National Academy of Sciences (2022). DOI: 10.1073/pnas.2116730119