March 25, 2022 feature
Study unveils new clonal relations in the mouse brain

The human brain and the brain of other mammals contain numerous populations of specialized cells with unique functions, molecular structures and characteristics. These cells originate from a thin layer of neuroepithelial progenitor cells, cells that can divide themselves into specific populations of neurons and glial cells.
In recent years, technological advances have allowed neuroscientists to study the diverse cell populations in the brain more in depth. While this has shed light on the function of some cell populations and their molecular composition, the relationship between mature cell populations and progenitor cells is still poorly understood.
Researchers at Karolinska Institute, KTH Royal Institute of Technology and Stockholm University have recently carried out a study aimed at better understanding the clonal relations between cells in the mouse brain. Their findings, published in Nature Neuroscience, were collected using a new approach they developed that combines single-cell and spatial transcriptomics with clonal barcoding, two different methods used to conduct neuroscience studies.
"Our lab studies the potential of neural stem cells to generate a wide variety of cell types, which is important to understand normal brain development and could be exploited to regenerate lost cells in neurological diseases," Michael Ratz, one of the researchers who carried out the study, told Medical Xpress.
To better understand the potential of stem cells as generators of different cell populations, the researchers used an approach known as "fate mapping" or "clonal tracking." This is a powerful technique that allows scientists to identify the "progeny" of a single cell and to reconstruct its developmental history (i.e., ancestry).
"These methods have led to fundamental insights about tissue development and have been used for many years, but they are limited in their ability to study many cells at the same time due to their reliance on microscopy which can only distinguish a few colors (usually <5) at once," Ratz said. "We wanted to establish an approach that matches the complexity of the nervous system, by converting the problem of fate mapping into a form that can be read out by modern high-throughput sequencing methods."
Past studies have found that cells in the mammalian brain can differ significantly in shape, function and spatial location. However, traditional methods to study the brain did not allow researchers to gather extensive information about individual cells.
An interesting metaphor that is sometimes used to describe this previous lack of insight about individual cells is the smoothie vs. fruit salad analogy. Essentially, previously neuroscientists were only able to observe large sections of brain tissues as a single, homogenous mixture (i.e., resembling a smoothie). This prevented them from learning more about specific cells in these sections of brain tissue (i.e., individual fruits inside the smoothie).
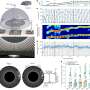
"In recent years, technological advances allowed us to look at individual cells at the mRNA level, to study the cellular composition of brain tissue and how cells that malfunction can lead to disease," Ratz explained. "One such method is single-cell transcriptomics, where individual cells are extracted from tissue and the sequences of thousands of mRNA molecules present in a single cell are sequenced while spatial transcriptomics achieves the same, but with intact tissue sections which preserves critical information about an individual cells' location in a tissue."
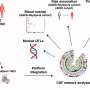
In their study, Ratz and his colleagues used genetic barcodes to characterize individual stem (progenitor) cells in the mouse brain. As these barcodes are inherited by daughter cells during brain development, they allowed them to study the clonal relationships between the cells and derive their phenotype profiles.
Subsequently, the researchers generated barcoding reagents and characterized the performance of these reagents in vivo (i.e., in the brain tissue of live mice). Their results could be very valuable for the neuroscientific community and could soon inform new studies investigating cell clonal relations more in depth. In the future, their experimental approach could also be used to conduct other studies examining specific cell populations or the relations between progenitor and daughter cells.
"We used this technology to discover unique clonal features of microglia, the brain's immune cells that play an important role in neurodegenerative disease, such as the large groups of cells an individual precursor cell forms and their widespread migration across large brain areas," Ratz added. "The tool we developed sheds new light on the molecular profiles underlying those cellular behaviors during normal development and could be adapted to study such microglia features in neurological diseases."
More information: Michael Ratz et al, Clonal relations in the mouse brain revealed by single-cell and spatial transcriptomics, Nature Neuroscience (2022). DOI: 10.1038/s41593-022-01011-x
© 2022 Science X Network