A new approach to therapy-resistant tumors targets a specific cell-death pathway
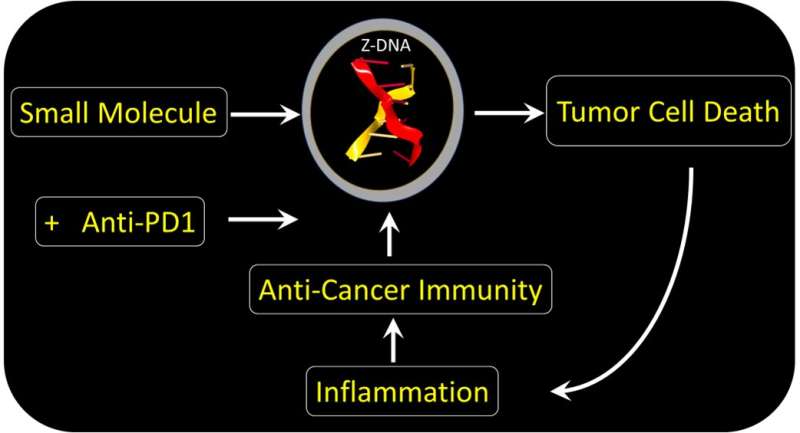
In a paper appearing in Nature today, an international group of scientists report a new way to kill hard-to-treat cancers. These tumors resist current immunotherapies, including those using Nobel Prize-winning checkpoint-blocking antibodies.
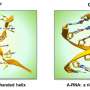
The approach exploits Z-DNA. Rather than twisting to the right like B-DNA, Z-DNA has a left-handed twist. One role for Z-DNA is to regulate the immune response to viruses. The response involves AADR1 and ZBP1, two proteins that specifically recognize Z-DNA. They do so through a Zα domain that binds to the Z-DNA structure with high affinity.
The Zα domain was originally discovered by Dr. Alan Herbert of InsideOutBio, a communicating author on the paper. The ADAR1 Zα domain turns off the autoimmune response, while the other ZBP1 Zα turns on pathways that kill virally infected cells, as previously shown by Dr. Sid Balachandran, the other communicating author on the paper. The interactions between ADAR1 and ZBP1 determine whether a tumor cell lives or dies.
Both Zα proteins are induced by interferon during inflammation. They are not usually present in normal cells. Both proteins are also expressed in tumors, especially in normal cells called fibroblasts that cancer cells force to support their growth. Normally, tumors rely on ADAR1 to suppress cell death pathways that would otherwise kill the tumor.
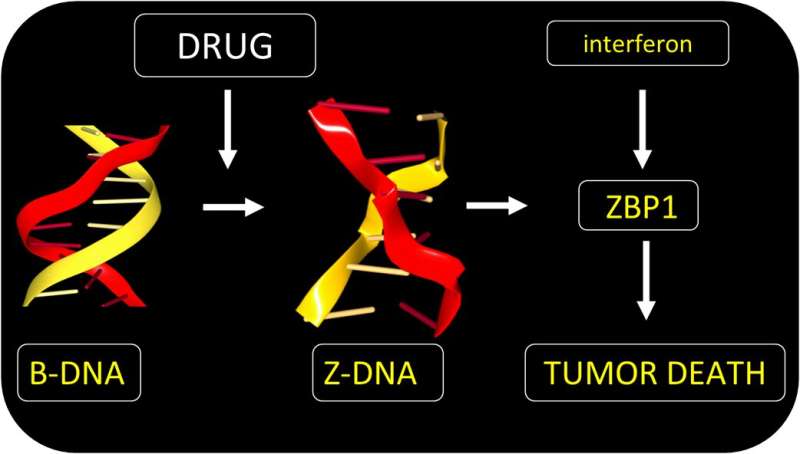
The team found a small molecule that could bypass ADAR1 suppression and directly activate tumor cell death by ZBP1. The drug acts regardless of the mutation that causes the cancer. The form of cell death induced is highly immunogenic. The response destroys the fibroblasts supporting the tumor growth. By doing so, the drug enhances the effectiveness of immunotherapy using the checkpoint-blocking antibody targeted at PD-1. The drug is a member of the curaxin family and was introduced to the clinic for another purpose. The compound has proven safe in Phase I trials, but still requires further research to confirm its clinical use in conjunction with anti-PD1 provides a benefit in the treatment of cancers.
Dr. Herbert says that "this outcome is the work of a highly collaborative team. It is a nice milestone in our understanding of how alternative DNA conformations, like Z-DNA, play an important role in human biology. The paper shows how basic research can lead to new and unexpected therapies. The process has taken a long time, starting with the initial discovery of the Zα domain and then to the identification of Zα DNA variants that cause genetic diseases in humans. These discoveries validated a biological role for Z-DNA. The work has now led us to a new therapeutic approach for the treatment of cancer. It is a pleasing turnaround, given that the initial Z-DNA discoveries were widely dismissed by the scientific community as of little biological importance and further work was not ranked as worthy of funding by the National Institutes of Health peer review panels."
Z-DNA is formed in normal cells by sequences called flipons that under physiological conditions can reversibly flip to the left-handed conformation. The structure was discovered by accident in the first ever synthetic crystal ever solved by X-ray crystallography. Other classes of flipons exist and are also highly likely to play important roles in biology. Flipons are highly dynamic structures that have been hard to study as it is challenging to determine their exact conformation inside cells. Dr. Herbert says that "it is similar to other highly dynamic systems in physics where you can only find the flipon conformation by making a direct measurement, but only if the act of measurement does not bias the result."
More information: Siddharth Balachandran, ADAR1 masks the cancer immunotherapeutic promise of ZBP1-driven necroptosis, Nature (2022). DOI: 10.1038/s41586-022-04753-7. www.nature.com/articles/s41586-022-04753-7