Researchers identify pathway that regulates angiogenesis in tumors

The formation of new blood vessels, also known as angiogenesis, is critical to the development and progression of cancer. Angiogenesis is controlled by many different protein messengers that can either stimulate or inhibit vessel growth. The protein YAP1 has been suggested to play a role in angiogenesis, but the molecular mechanisms of this process are not completely understood. In a new article published in Cancer Research Communications, Moffitt Cancer Center researchers describe the signaling pathways that regulate YAP1 and how the protein contributes to angiogenesis under normal and low oxygen conditions.
Cancer is associated with abnormal blood vessel growth, resulting in limited nutrients and oxygen in the tumor environment because of poor transport. Low oxygen levels, also known as hypoxia, limit the ability of cancer cells to survive. Therefore, in hypoxic conditions tumor cells trigger angiogenesis by upregulating multiple signaling pathways, such as YAP1, that regulate blood vessel growth, patterning and function.
In a series of laboratory experiments, Moffitt researchers found that YAP1 controls the expression of several genes involved in the formation of new blood vessels, confirming the importance of YAP1 to angiogenesis. Next, the team focused their attention on delineating the molecular pathways that regulate YAP1 under normal and hypoxic oxygen conditions and the downstream pathways that YAP1 controls during angiogenesis.
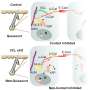
The researchers discovered that YAP1 activity and its cellular localization are highly dependent on the levels of oxygen in the tumor environment. Under normal oxygen conditions, YAP1 is located in the cytoplasm and the nucleus of the cell. However, under hypoxic conditions, the majority of YAP1 is found in the nucleus.
The team next identified the key molecular events that led to this transition. Under normal oxygen conditions, YAP1 is altered by the protein PHD2, which adds hydroxyl chemical modifications to YAP1. This results in interactions between YAP1 and the protein VHL, which targets YAP for protein degradation. Alternatively, under hypoxic conditions, YAP1 is unable to interact with PHD2 and becomes more stable, enabling it to interact with the proangiogenic protein HIF1α, leading to increased expression of proteins involved in blood vessel growth.
Importantly, the Moffitt researchers confirmed a clinical connection between YAP1 and tumor development. They discovered that nuclear expression of YAP1 was higher in samples from renal cell carcinoma patients than normal kidney tissue. Additionally, interactions between YAP1 and the protein HIF1α were higher in tumor tissue than normal tissue.
These combined data demonstrate the importance of YAP1 to angiogenesis and suggest that YAP1 may be a potential target for anticancer drugs, particularly in renal cell carcinoma.
"We have shown that YAP1 is regulated by a novel post-translational modification that involves proline hydroxylation at a specific region, which is sensitive to oxygen levels. These findings, along with the role of YAP1 in neoangiogenesis, opens new avenues to combat various cancers that are driven by YAP1, as well as tumors like renal cell carcinoma, which show loss of VHL along with elevated YAP1 levels," explained Srikumar Chellappan, Ph.D., study author and chair of the Department of Tumor Biology at Moffitt.
More information: Namrata Bora-Singhal et al, A novel PHD2/VHL- mediated regulation of YAP1 contributes to VEGF expression and angiogenesis, Cancer Research Communications (2022). DOI: 10.1158/2767-9764.CRC-21-0084