How common genetic alterations cause kidney cancer
An international group of researchers investigated why mutations often associated with renal cancer result specifically in renal cancer, instead of other cancer types.
Typically, gene mutations occurring in cancers are associated with only certain subtypes of cancer. The reasons for this have remained largely unclear.
According to the study published in Nature, the ability of mutations associated with renal cancer to drive the onset of cancer is based on proteins known as transcription factors, which regulate the normal functions of healthy renal cells.
"The study focuses on renal cancer, but the same principle may be applicable to cancers more generally. Cancer is often an incurable disease, and new therapies are needed. We identified factors whose inhibition slowed down the growth of renal cancer in experimental models. These molecules are potential drug targets," says Sakari Vanharanta, the project's principal investigator at the University of Helsinki.
A common gene mutation increases the risk of renal cancer
The mechanisms examined in the study concern clear cell renal cell carcinoma, the most common subtype of renal cancer. Most Europeans carry a genetic variant that increases the risk of developing this disease.
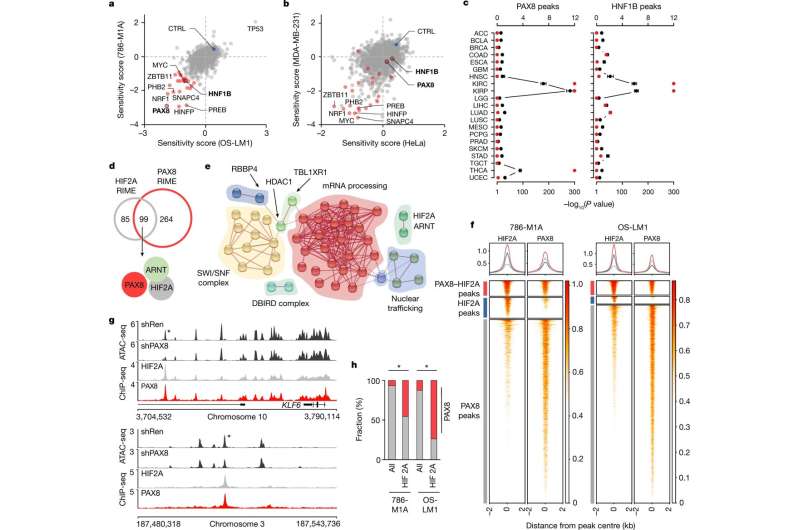
In the study, methods of experimental cancer biology were utilized in conjunction with targeted patient specimen analysis. The researchers tested whether transcription factors that control the tissue-specific functions of normal renal cells are also needed for the growth of renal cancer.
The results indicate that the ability of mutations associated with renal cancer to promote tumor formation was dependent on transcription factors that are particularly active in normal renal cells. When renal lineage-specific transcription factors were experimentally inactivated, cancer mutations were no longer able to activate genes relevant to tumor growth.
"The role of of genetic alterations, or mutations, in carcinogenesis is well established, but what we often don't know is how cancer mutations cause cancer. This makes it difficult to exploit the genetic information in cancer therapy. Understanding the functional consequences of cancer-related mutations is therefore important, and it can, as it has already in some cases, help scientists develop new cancer drugs," Vanharanta says.
More information: Saroor A. Patel et al, The renal lineage factor PAX8 controls oncogenic signalling in kidney cancer, Nature (2022). DOI: 10.1038/s41586-022-04809-8