How the thymus trains T cells to fight infections
T cells are a special class of white blood cells that patrol the body and attack infected or foreign tissue. They learn to distinguish friendly proteins from dangerous ones in an organ called the thymus. However, when T cells mistakenly identify healthy proteins as foreign, it can lead to autoimmune disorders such as multiple sclerosis or diabetes.
New work from Cold Spring Harbor Laboratory Fellow Hannah Meyer describes how the human thymus generates the list of friendly proteins that T cells should not attack. Her team identified, for the first time, the RNA molecules used to generate this list that protects healthy tissue from T cells. Their discovery may help identify key differences between effective and flawed immune systems, and lead to improved autoimmune disorder treatments.
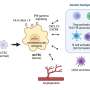
To attack the right targets, T cells need a thorough education about the proteins they might encounter in the human body. The human body can produce around 20,000 different types of proteins. Before T cells leave the thymus to fight infections, they must be trained to recognize all of these friendly proteins. This means the thymus has to make all 20,000 proteins. No other organ in the human body makes every possible protein. They only make the proteins needed for their specific organ function. "Usually, it is very strictly regulated which proteins are made in which cells and tissues. Thymus cells make all of them to ensure the immune system functions properly," Meyer says.
Meyer focuses on the thymus for its role in not just preventing autoimmune diseases but for also fighting infections and cancer. Meyer hopes these diseases can be better understood and treated by further exploring the proteins made in the thymus. Meyer's new list is available to other researchers in an interactive online database. The research is published in Nature Communications.
More information: Jason A. Carter et al, Transcriptomic diversity in human medullary thymic epithelial cells, Nature Communications (2022). DOI: 10.1038/s41467-022-31750-1
Interactive database: epitopediversity.cshl.edu/