Study illuminates precancerous 'clonal outgrowth' in blood cells
A common, spontaneous mutation in blood stem cells, which has been linked to higher risks of blood cancer and cardiovascular disease, may promote these diseases by altering the stem cells' programming of gene activity and the mix of blood cells they produce, according to a study co-led by investigators at Weill Cornell Medicine, NewYork-Presbyterian, the New York Genome Center, Harvard Medical School and Dana-Farber Cancer Institute.
The blood stem cell mutation, known as DNMT3A R882, leads to the growth of a large population, or "clonal outgrowth," of circulating blood cells that also contain this mutation. In general, such mutant outgrowths become increasingly common with age, and are thought to represent a very early, pre-malignant stage of cancer development.
However, the molecular details of how they arise have been hard to pin down, because the mutant cells broadly look and function the same as normal cells. In the study, which appears in Nature Genetics, the researchers surmounted this challenge to illuminate the effects of R882 mutations in DNMT3A, the most commonly mutated gene in blood cells.
"These findings help us understand how these mutated cells outgrow normal cells, and pave the way for possible future interventions targeting these cells to prevent cancers and other clonal outgrowth-related conditions," said study senior author Dr. Dan Landau, an oncologist at NewYork-Presbyterian/Weill Cornell Medical Center.
The study was a collaboration between Dr. Landau's laboratory and the laboratory of Dr. Irene Ghobrial, professor of medicine at Harvard Medical School and the Dana Farber Cancer Institute. Dr. Ghobrial's team supplied samples of blood stem cells from the marrow of patients in remission from multiple myeloma—patients in which, they have found, blood cell clonal outgrowths are relatively common.
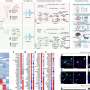
Dr. Landau's team evaluated more than 6,000 cells from the patients, using "single-cell multi-omics" techniques to detect the DNMT3A R882 mutation, and to map gene activity and chemical marks on DNA called methylations, programming marks that switch off nearby genes. In this way, they recorded in unprecedented detail how the mutation-containing blood stem cells differed from their normal counterparts.
The researchers found, for example, that the mutant stem cells' production of mature blood cells was skewed towards red blood cells and the cells that make blood-clotting platelets—providing potential rationales underlying the higher risk of cardiovascular disease in patients with clonal outgrowths in their blood.
The gene DNMT3A normally encodes an enzyme called a methyltransferase, which helps place methylations on DNA. The researchers found that the mutation's disruption of normal methylation led to a lack of these "off switches" across the genome and the abnormal activation of key genes. The latter included inflammation-driving genes and cancer-associated growth genes—all consistent with a growth and survival advantage for the mutant cells, and a higher risk of their progression to cancer.
"Our hope is that by uncovering molecular signatures like these we'll be able to target these clonal outgrowths and prevent cancer development in people who are still healthy," said study co-first-author Dr. Anna Nam, assistant professor of pathology and laboratory medicine in the Department of Pathology and Laboratory Medicine and a member of the Meyer Cancer Center at Weill Cornell Medicine, and a pathologist at NewYork-Presbyterian/Weill Cornell Medical Center.
The researchers plan to do further studies of clonal outgrowths resulting from other mutations. They are also developing their multi-omics techniques to increase the speed and scale of these studies.
"We should soon be able to do studies of many more cells at a time, giving us a more complete picture of what is going on," said co-first author Neville Dusaj, a Tri-institutional MD-Ph.D. Program student in the Landau laboratory.
More information: Irene Ghobrial, Single-cell multi-omics of human clonal hematopoiesis reveals that DNMT3A R882 mutations perturb early progenitor states through selective hypomethylation, Nature Genetics (2022). DOI: 10.1038/s41588-022-01179-9