Seeking microscopic clues to beating deadly brain tumors
A critical new pathway to treating an aggressive brain tumor might be found in the complex diversity within the tumor tissue, according to a new paper by scientists from the Hackensack Meridian Center for Discovery and Innovation (CDI).
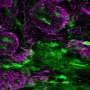
The CDI laboratory deeply analyzed tumor tissue using an advanced mass spectrometry with special focus on lipids, a class of molecules that includes fats, according to the new paper, in the journal Scientific Reports.
"Lipid ions presented here lay the foundation for future studies that are required to understand their interconnecting signaling pathways in relation to cell function, tumor progression, and resistance to therapy," according to the paper. "Understanding their functional relevance is essential for the identification of new therapeutics based on lipid pathway targets."
Glioblastoma is one of the most difficult cancers, let alone diseases, to treat. The brain tumor presents a median survival rate of just 12 to 15 months.
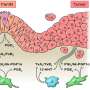
The cancer is especially hard to beat, since it presents so heterogeneously—with different tumor cell subtypes within the same tumor, which can all respond differently to therapy. The cancer also has a tendency to create aberrant small blood vessel networks quickly, helping it spread quickly—and making it particularly hard to defeat through traditional treatment pathways.
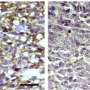
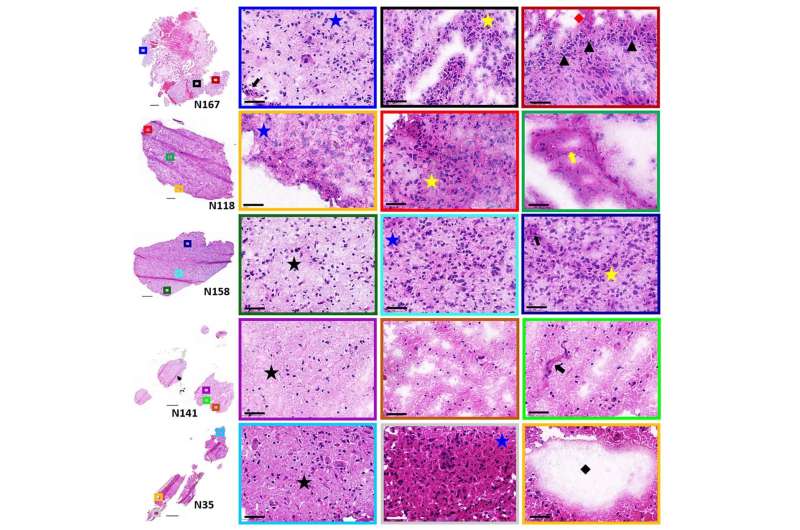
The CDI scientists assessed five human samples of brain tumor tissue. The team looked at a series of different lipids, in different sections of the tumor and the surrounding environment. They found a series of possible treatment candidates, according to the publication.
The corresponding author, Claire Carter, Ph.D., is an expert in a special type of mass spectrometry imaging called "Matrix-Assisted Laser Desorption/Ionization," or MALDI. Carter led the work, and it is part of ongoing looks into the tumor microenvironment (TME) in glioblastomas and other deadly pediatric tumors that she is taking with colleagues including Dr. Derek Hanson, Director, Pediatric Neuro-Oncology at Hackensack University Medical Center .
"In conclusion, high resolution MALDI MSI identified a number of lipids that differentiate tumor and endothelial cell subpopulations within human glioblastoma samples," the authors write. "The heterogenous distributions… within these cell population further highlight the complexity of the glioblastoma TME."
The scientists hypothesize that a multi-pronged approach may fare best against the stubborn cancer.
"Targeting several of these lipids and their signaling pathways simultaneously, however, may improve clinical outcome," they write.
More information: Kelly C. O'Neill et al, Mass spectrometry imaging discriminates glioblastoma tumor cell subpopulations and different microvascular formations based on their lipid profiles, Scientific Reports (2022). DOI: 10.1038/s41598-022-22093-4