November 4, 2022 report
Using single-cell sequencing to reveal processes involved in ovarian and breast cancers
A team of researchers from Weill Cornell Medicine, the British Columbia Cancer Research Centre and Memorial Sloan Kettering Cancer Center, working with a group from the IMAXT Consortium, has used single-cell sequencing to reveal breast and ovarian cancer-related mutational processes.
In their paper published in the journal Nature, the group describes how they conducted single-cell genome sequencing on thousands of mammary tissue cells and compared what they found with sequencing they performed on thousands of ovarian and breast tumor cell samples.
The researchers began their study after noting that cell-to-cell copy number alterations that can trigger genomic instability in various kinds of cancer have not been well addressed by the research community. They also noted that the means by which such alterations can lead to variations in phenotypes in the evolution of different kinds of cancers has also not been adequately studied. To rectify the situation, the team embarked on an ambitious sequencing effort focused most directly on mutational processes related to ovarian and breast cancers.
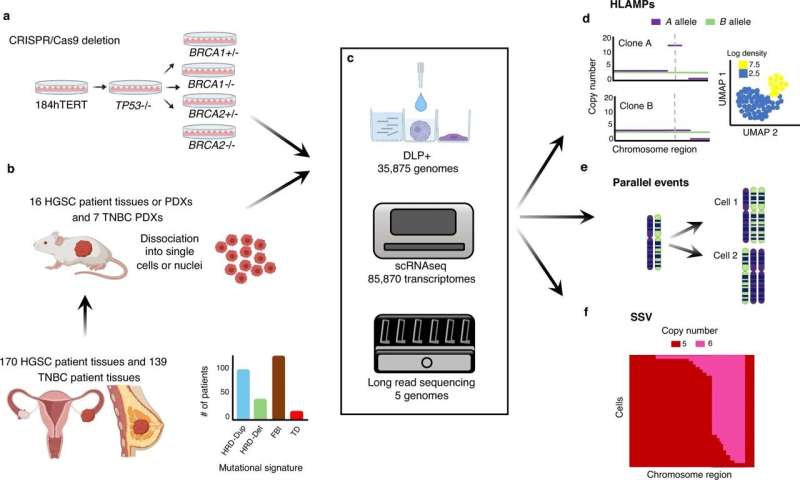
The work by the team was two-pronged. One effort involved conducting single-cell genome sequencing of 13,800 mammary epithelial cells collected from women who may or may not have had mutations in tumor protein p53 or breast cancer type 1 or 2 susceptibility proteins—resulting in homologous recombination deficiency. They followed that up by looking at haplotype patterns and single-cell structural variants to find mutual processes.
The other effort involved conducting single-cell genome sequencing on 22,057 advanced ovarian or breast cancer tumor cells. They then compared patterns they found in the first effort, which they describe as foreground events, with those found in the second effort.
In their comparisons, the researchers found what they describe as cell variations in copy number and structural alterations that were widespread in triple-negative breast cancers and in high-grade serious carcinoma, which were seen in distinctive instability processes. They further suggest that such alterations indicate an influence on expression levels of hundreds of genes.
The researchers also found other differences in the cells they studied that might help shed light on the many processes that lead to the development of ovarian and breast cancers.
More information: Tyler Funnell et al, Single-cell genomic variation induced by mutational processes in cancer, Nature (2022). DOI: 10.1038/s41586-022-05249-0
© 2022 Science X Network