Whole-genome sequencing offers a more comprehensive prenatal test
BGI-Research and the Maternal and Child Health Hospital of Hubei Province (MCHH) have published whole-genome sequencing research results in npj Genomic Medicine. These results have revealed the genomic architecture of fetal central nervous system (CNS) anomalies systematically in a large cohort for the first time and share how whole-genome sequencing may enhance detection of fetal CNS anomalies.
Fetal central nervous system (CNS) anomalies are among the most common congenital abnormalities with the incidence of neural tube defects (NTDs) alone reaching 18.6 per 10,000. For every parent who hopes to have a healthy baby, prenatal tests provide critical information for clinical decisions.
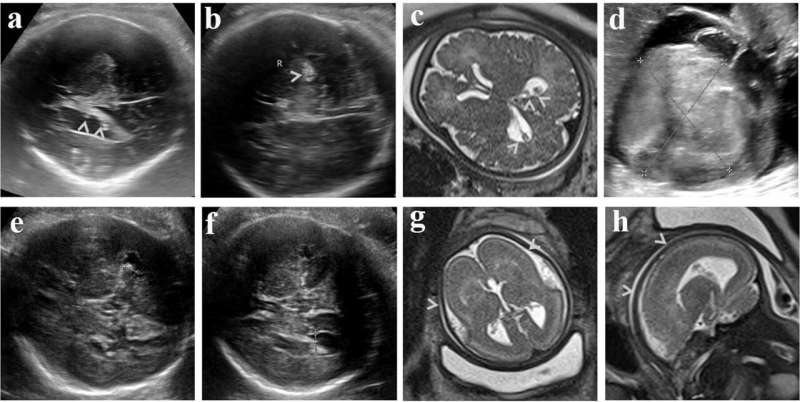
Currently, ultrasonography is the most important modality for evaluating fetal CNS defects. Diagnosing fetal CNS anomalies through medical imaging is still very challenging at this stage because the fetus and its brain are constantly developing.
Although hundreds of genetic conditions involving the CNS have been previously studied, there is still a lack of research on the traits they exhibit in imaging tests. In addition, there has been a lack of large cohort genomics studies for fetal CNS abnormalities.
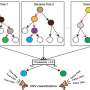
This study was divided into two phases, first using low-depth whole-genome sequencing to analyze aneuploidy and copy number variants, and then continuing with 40X whole-genome sequencing on low-depth whole-genome sequencing negative samples to further analyze single nucleotide variants and small fragment copy number variants to systematically resolve genes associated with fetal CNS abnormalities and to demonstrate the role of whole-genome sequencing in prenatal disease diagnosis and its potential as a clinical diagnostic tool.
From 2015 to 2017, the research team collected 162 clinical samples related to central nervous system anomalies. A total of 18 aneuploidy variants, 21 copy number variants, three small fragment copy number variants and 26 single nucleotide variants, including 15 novel mutations, were identified as pathogenic or potentially pathogenic in this study. Through statistical analysis of these variants, 62 cases of CNS anomalies were effectively diagnosed, with a detection rate of 38.3%. Among these cases, the detection rate of microcephaly and total forebrain anomalies was over 70%.
Of the 29 key genes containing diagnostic variants detected in this study, 5 were present in more than one sample. These diagnostic variants were detected in 70.8% of fetuses with both CNS and non-CNS anomalies, compared to 24.6% of fetuses with only CNS anomalies.
It was also found that whole-genome sequencing is a more comprehensive test than other tests currently used in clinical practice. Low-depth whole-genome sequencing assays had similar detection rates for microscopic copy number variants as karyotyping, and detected an additional 2.3% of submicroscopic pathogenic copy number variants in samples that did not contain microscopic copy number variants.
In addition, the fetal anomalies detection rate of 40X whole-genome sequencing assay further increased by 18.9% compared with the low-depth whole-genome sequencing assay, and the overall detection rate was higher than that of whole-exome sequencing.
Considering the clinical application, this study also analyzed the cost and time of whole-genome sequencing. On the BGI platform, the sequencing costs of whole genome sequencing are lower than that of combined chromosome microarray and exome sequencing. In addition, the analysis time is reduced to a level similar to that of exome sequencing, and even shorter than that of combined chromosome microarray analysis and exome sequencing.
Although whole-genome sequencing assays showed obvious advantages in this study, the actual clinical significance of some of the variants detected in this study is currently difficult to determine due to the lack of database constraints at this stage. In the future, the application of whole-genome sequencing in prenatal testing can be further improved if more samples can be analyzed and the analysis can be extended to each anatomical system.
BGI will continue to take advantage of multi-omics technologies to carry out in-depth research related to maternal and infant health, to further support the development of clinical testing technologies and bring more possibilities for prenatal testing of genetic diseases.
More information: Ying Yang et al, Genomic architecture of fetal central nervous system anomalies using whole-genome sequencing, npj Genomic Medicine (2022). DOI: 10.1038/s41525-022-00301-4