Researchers map activity of inherited gene variants linked to prostate cancer

UT Southwestern researchers have identified the molecular function of 87 inherited genetic variants that affect the risk of prostate cancer, and the majority appear to control the activity of genes located far away from the risk variants themselves. The findings, published in Cancer Discovery, could lead to better ways to assess cancer risk and new targets for anti-cancer drugs, the study authors say.
"Traditionally, we think of regulatory elements in the genome affecting neighboring genes," said study leader Ram Mani, Ph.D., Assistant Professor of Pathology and Urology at UTSW and a member of the Harold C. Simmons Comprehensive Cancer Center. "But these risk variants, or risk alleles, can act like a light switch. The light is on the ceiling, but the switch is on the wall on the other side of the room."
The findings also indicate that an individual risk allele can regulate multiple genes, he said.
Prostate cancer is one of the most heritable cancers, Dr. Mani explained. Researchers have identified at least 185 common prostate cancer risk alleles, or snippets of DNA, passed down through families of European descent. But because the vast majority of these alleles are located in DNA's noncoding regions—which don't contain genes and therefore don't produce proteins—how they affect prostate cancer risk has largely been a mystery.
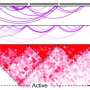
Previous research has suggested that they may control gene regulation, but which genes are affected and how their activity is altered has been unknown. To answer these questions, Dr. Mani and his colleagues used several approaches to identify which genes serve as targets for the risk alleles. A three-dimensional mapping technique using data from 565 prostate cancer tumors showed that 87 of these risk alleles affected the activity of hundreds of genes.
Although malignant tumors typically arise in the prostate's epithelial cells, researchers found that the affected genes were often in other tissue types, including stromal cells and smooth muscle cells that support the epithelial cells. Most of the risk alleles appeared to alter the activity of these genes, which produced proteins known to be involved in molecular pathways for development, apoptosis (programmed cell death), and metabolism, among other cellular processes.
Dr. Mani said some alleles had opposing activity on the multiple genes they control. For example, one allele known as rs8102476 simultaneously increased the activity of one gene while decreasing the activity of a neighboring gene. The risk alleles also had significant interaction with genes that acquired nonheritable mutations associated with prostate cancer; these interactions appeared to predict how aggressive a patient's disease became.
Together, Dr. Mani said, these findings could lead to better risk models for patients as well as new prostate cancer treatments. Further association studies are urgently needed to identify risk alleles in populations of non-European descent so that risk models and treatments can be customized to a broader patient base, he added.
"African American men have an elevated risk of prostate cancer, especially aggressive disease," Dr. Mani said. "There's a critical need for more genomewide association studies in this and other populations in which there's an established health disparity to understand the nature of their risk and how to decrease it."
More information: Jiapei Yuan et al, Prostate Cancer Transcriptomic Regulation by the Interplay of Germline Risk Alleles, Somatic Mutations, and 3D Genomic Architecture, Cancer Discovery (2022). DOI: 10.1158/2159-8290.CD-22-0027