This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
Advanced computing indicates current antibodies effective against SARS-CoV-2 variant XBB.1.5
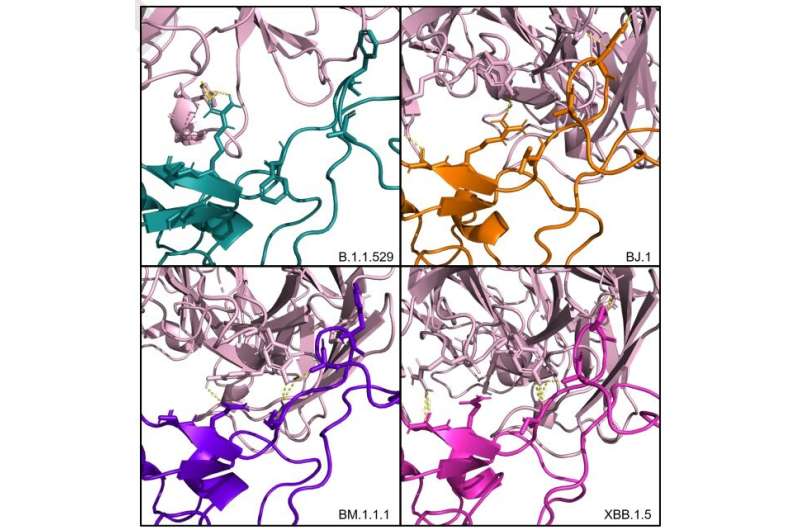
A team at UNC Charlotte's Center for Computational Intelligence to Predict Health and Environmental Risks (CIPHER) and Tuple, a Charlotte-based genomics consulting firm, has used artificial intelligence to rapidly assess the public health implications of the newly emergent SARS-CoV-2 XBB.1.5 variant. Results from simulations run by the team indicate the antibodies currently in our arsenal are effective to neutralize SARS-CoV-2 XBB.1.5. This work is available on the bioRxiv pre-print server
The variant has recently increased its share of COVID-19 infections caused in the United States, creating concern about a potential health care crisis. However, increased transmission ability of the variant does not mean it is not treatable.
"The CIPHER and Tuple team have tied several technologies together to provide rapid health intelligence, which is a dose of good news. It is becoming more clear that with new sciences and hard work we can address emergent diseases," said Daniel Janies, the co-director of the CIPHER center and Carol Grotnes Belk Distinguished Professor of Bioinformatics and Genomics.
The team used machine learning applications to simulate protein structure and antibody binding. That way, they tested a variety of antibodies including those induced by the omicron booster vaccine and others used in therapies.
These results are in contrast to those found by the team when the original omicron variant emerged over a year ago. In 2021, the team correctly predicted the original vaccine and antibody arsenal had reduced efficacy against omicron.
As this team works in computational simulations on digital representations of the variants and the antibodies, the work is very quick. The team uses protein structures from the program AlphaFold2, then a program called HADDOCK to dock the antibodies to the variants' receptor binding domain in the Spike protein of SARS-CoV-2 XBB.1.5 and related viruses. The assessment is based on physics models to test the quality of docking and predict maintained or weakened efficacy of the antibodies against the viruses.
The computational work precedes, by weeks, the structural and binding experiments that must be done in a traditional chemistry laboratory. CIPHER researchers leverage new tools and UNC Charlotte's advanced computing capabilities to provide a valuable early indication of what health officials can expect from the rise of a new variant as well as the therapeutics and vaccines that are effective in response.
The peer reviewed omicron paper, "Predictions of the SARS-CoV-2 Omicron Variant (B.1.1.529) Spike Protein Receptor-Binding Domain Structure and Neutralizing Antibody Interactions," which used the same methodology and was published last year in Frontiers in Virology can be found here.
More information: Colby T. Ford et al, Positing changes in neutralizing antibody activity for SARS-CoV-2 XBB.1.5 using in silico protein modeling, bioRxiv (2023). DOI: 10.1101/2023.02.10.528025