This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Genome analysis just got personal

Life on Earth comes in a beautiful assortment of different shapes, sizes and colors, thanks to genetic mutations. Some mutations are beneficial, some are perilous, and some don't do much of anything. Every person has around 4.5 million genetic variations. But are those variants helpful or hurtful? Geneticists have been trying to find the answer for half a century. Their biggest obstacle nowadays? Standard human genome sequence reference data.
The original human genome sequence is a combination of 13 individual donors, with little to no ethnic diversity between them. A more personal genome sequence is needed to understand what mutations cause diseases in a single individual. Cold Spring Harbor Laboratory Professor Thomas Gingeras and Yale University Professor Mark Gerstein are leading an international, multi-institutional effort to meet this need.
Gingeras says, "It is very clear, for a long time, that the ideal would be to get everybody's genome sequence and do the analysis of cause and effect [on] the variations as the basis of diagnoses and their treatment. This is where medicine is going. And this is an attempt to provide a paradigm for doing that."
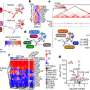
They've now sequenced four people's genomes and tracked the mutations in each of them, along with their genetic consequences. The team created the world's largest catalog of genetic mutations called allele-specific variants. Using this catalog—EN-TEx—they built an algorithm to predict how the variants affect tissues and a person's risk for developing certain diseases. The catalog and algorithm provide an unprecedented tool for personalized medicine. The work is published in the journal Cell.
"We mapped over a million allele-specific variants in each of the four sequenced individuals," Gingeras says. "Our findings indicate that parts of the genome, called cis-regulatory elements, can be particularly sensitive to these genetic variants. Overall, EN-TEx provides rich data and models for more accurate personal genomics."
For scientists, one of the key features of this new approach is the ability to study the effects of genetic mutations in tissues that are difficult to obtain without surgery. For example, if someone had a heart or brain condition, performing genomic analysis on those tissues would be challenging unless there was a clinical need to operate. But with this new method, the analysis could be done using a person's blood as a "surrogate."
Gingeras hopes his work will bring us a step closer to personalized medicine. Collecting and digging through thousands of genomic data points is a formidable task. Gingeras' "blueprint" could make it much more manageable.
More information: Mark Gerstein, The EN-TEx resource of multi-tissue personal epigenomes & variant-impact models, Cell (2023). DOI: 10.1016/j.cell.2023.02.018. www.cell.com/cell/fulltext/S0092-8674(23)00161-7