This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Refining your search: A team approach to identifying patient cohorts using data from the electronic health record

Finding the right patients for clinical trials can be a struggle and require much time and effort. Clinical researchers have wondered whether screening the electronic health record (EHR) could improve the process. Could it provide enough information to determine whether a patient is a good fit for a study?
Clinical research tools like i2b2 promise to make such EHR searches easier for clinicians. Much like someone can develop search terms and syntax for a search for library materials, clinicians can use i2b2 to develop search queries for EHR data to find patients for clinical trials. But what if their queries are not accurately identifying the patients who would be a good fit for their studies or are missing eligible patients? How can these queries be improved?
A recent article by MUSC researchers in the Journal of the American Medical Informatics Association sought to answer that question. The study was led by Alexander Alekseyenko, Ph.D., and Bashir Hamidi of the MUSC Biomedical Informatics Center.
Although clinicians have begun to use tools like i2b2, they tend to create simple queries that may not be precise enough to identify patients who fit a trial's criteria. Considerable finesse is required when designing these queries, as some criteria are not so easily defined in the EHR.
For example, a diagnostic code in the EHR may not be enough to ensure that a patient has the disease of interest to the trial. Clinicians may use codes somewhat differently.
"Clinicians might actually see patients slightly differently and sometimes code them differently," said Kit Simpson, DrPH, Distinguished University Professor in the Department of Health Care Leadership and Management at MUSC and a co-author of the study. "As a result, researchers might miss a large number of people who would have been eligible for the trial."
Likewise, a clinician might assign a code based on his or her assessment of symptoms without validating the diagnosis through testing.
"Suppose we have a child who has frequent breathing problems," said Alekseyenko. "You come to the doctor, and he or she says, 'This child has a reactive airway disease' and enters it as a diagnosis based purely on symptoms and not any tests."
If a researcher is looking for patients with reactive airway disease, that child would come up in the search, even though the diagnosis has not been verified by a test.
Further specifying the criteria could improve results. For example, one workaround would be to require several mentions of the same diagnostic code in the health record within a specific time frame. If that diagnostic code comes up several times for that child, that would make it more likely that the child has the disease.
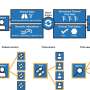
Like a library user turning to a research librarian for help with a search, clinicians can develop more accurate and specific "e-phenotypes"– or sets of EHR criteria that identify patients as eligible for studies—by working with experts in the field. These include biomedical informaticists and data architects, who know how to build the search queries, and the "honest brokers," who know what information is available in an institution's EHR and where to find it. An honest broker is a neutral intermediary who acts on behalf of all parties, collecting and providing de-identified information to research teams in an impartial manner.
"It's just like using available statistical programs. I can plug numbers into them and just keep hitting buttons, and it'll give me a result, but it may not mean anything," said Patrick Flume, M.D., co-director of the South Carolina Clinical & Translational Research (SCTR) Institute and a co-author of the study. "If you really want to make sure you get meaningful results, you need to work with people who know how to use the tool properly."
The study asked 21 clinical trial leaders from a wide variety of specialties to work with an informaticist and honest broker to define a phenotype and build a search query for the EHR. Each search query was then used to identify 20 patients who met study criteria. These same clinical trial leaders were then asked to go through the patients for their disease of interest to determine how well they matched study criteria.
Results were mixed, with matching being better for some e-phenotypes, such as infection, neonatal conditions and cancer, than others, including psychiatric, gastrointestinal and pulmonary disease. Better matching was also seen for patients who received inpatient rather than outpatient care, as more data are collected in the EHR during hospitalization.
Interestingly, clinician confidence did not correlate with better matching, though better-specified phenotypes did.
The study demonstrates that it is possible to use e-phenotypes to identify patients who match clinical trial criteria but also that more work remains to be done to refine those phenotypes to return more accurate results. It also suggests that a prerequisite of success is a team approach drawing on clinical, informatics and database experts.
In addition, continuing to refine e-phenotypes to improve matching is crucial to unlocking their potential. Further, better-refined e-phenotypes could make clinical trial enrollment much more efficient.
"Study coordinators may be given lists of patients right now that they have to manually sort through and read their charts one at a time to see if they're eligible," said Hamidi. "But if we are able to create these precise, accurate phenotypes, the entire process could be much more efficient because they could be sure that a particular patient falls into the group that they're interested in."
The ability to identify eligible patients quickly could be especially important for quick-moving diseases like sepsis or time-sensitive clinical trials, such as those based in the intensive care unit.
"Going bed to bed and reading charts in the ICU is time consuming," said Flume. "If you had a method to filter the number of patients down to the few who truly are eligible for the trial, then you could be far more efficient at screening these people for participation in a trial."
It is also important for efforts like MUSC's Living Biobank, which creates an Institutional Review Board (IRB)-recognized process for making unused clinical specimens available for research. The IRB reviews and monitors clinical research and has the right to approve, require modifications to or disallow research in accordance with Food and Drug Administration guidelines. E-phenotyping would make it easier to fulfill requests from IRB-approved studies for patient-derived specimens before they are discarded. Those specimens could be used to generate preliminary data or verify that patients meet clinical trial criteria. E-phenotypes could also be used to identify historical controls for small clinical studies. Finally, they could help to identify patients from all ethnic and racial backgrounds, making clinical trial participants more reflective of the community and of the affected patient population.
"Once we get a bunch of good e-phenotypes, we can actually use them to make sure that we have representation in underserved areas in South Carolina so that people who would not normally be asked in time get on the list and get contacted," said Simpson. "And that goes for minorities, for rural patients and for patients in medically underserved areas."
More information: Bashir Hamidi et al, Not all phenotypes are created equal: covariates of success in e-phenotype specification, Journal of the American Medical Informatics Association (2022). DOI: 10.1093/jamia/ocac157