This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Study sheds light on a mitochondrial disease
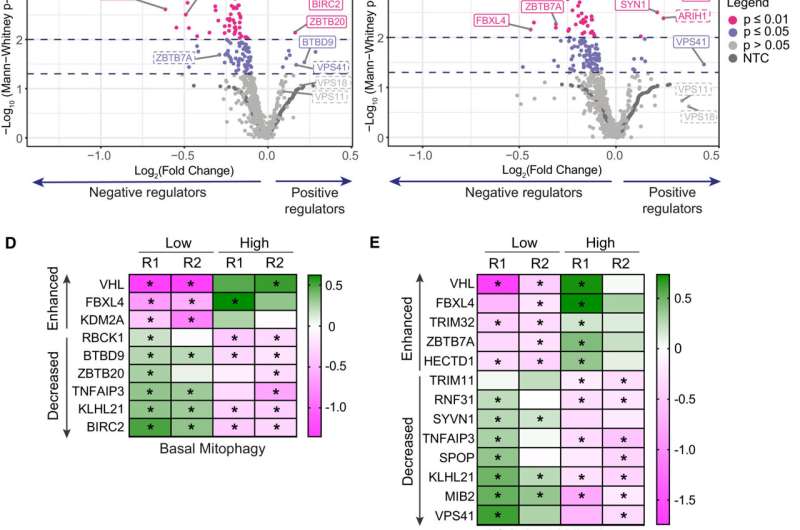
Scientists at the University of Liverpool have figured out how mutations in a gene called FBXL4 can lead to an excess of mitophagy—the disposal of mitochondria, the 'power stations' within nearly all human cells.
Defects in mitophagy have been linked to neurodegenerative ailments such as Parkinson's and Alzheimer's disease and many other pathophysiological conditions. However, too much mitophagy can be detrimental as well. In this study, a team of scientists led by Professor Sylvie Urbé and Professor Michael Clague and supported by Parkinson's UK, reveal how mutations in the FBXL4 gene lead to an excess of mitophagy.
The findings show that FBXL4 controls the levels of another protein called NIX, which can dive into the mitochondria and effectively act as a notice of condemnation.
Professor Sylvie Urbé said, "From time to time, mitochondria are decommissioned and new ones are made. The old or damaged mitochondria are delivered to the cell 'breakers yard' or lysosome in a process called mitophagy. Mitophagy is important in developmental pathways but also in suppressing neurodegeneration and other pathophysiological conditions."
"Sometimes mitophagy can be over-active and patients are left with too few mitochondria for the bodies energy needs and suffer from mitochondrial depletion syndrome. This is the case for patients with a mutation in a gene called FBXL4."
The work is published in The EMBO Journal alongside complementary findings from laboratories in China and Australia. The findings will help to open up new avenues for treating these mitophagy related diseases.
More information: Hannah Elcocks et al, FBXL4 ubiquitin ligase deficiency promotes mitophagy by elevating NIX levels, The EMBO Journal (2023). DOI: 10.15252/embj.2022112799