This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers invent powerful tool to gather data on immune response at single-cell level
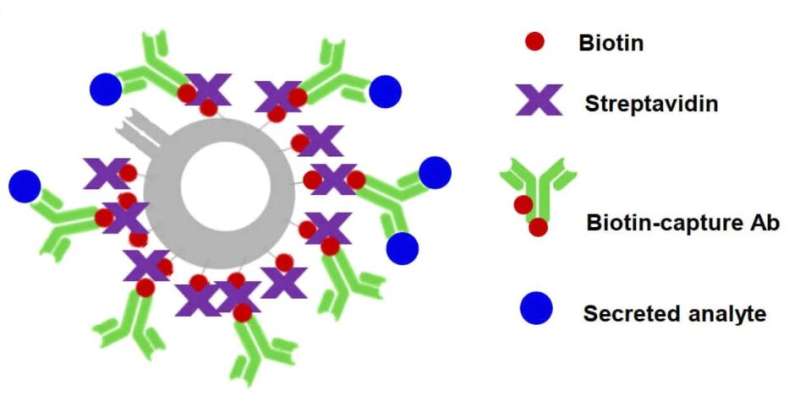
Cells interact with their surrounding environment by secreting proteins which act as messengers or signals for communicating with other cells. Capturing these elusive and minute secreted proteins, particularly those from our immune cells, and correlating them to the individual source cells can provide important insights into immune responses in patients with chronic diseases, such as cancer, autoimmune disorders, or infectious diseases, and accelerate the development of immunotherapies.
However, studying how unhealthy or healthy cells communicate, interact and coordinate with each other in response to stimuli or pathogens, remains a challenge for scientists.
To help fill the gap in correlating cell functions with their secreted proteins, a team of researchers from the National University of Singapore (NUS) led by Assistant Professor Cheow Lih Feng from the Department of Biomedical Engineering under the NUS College of Design and Engineering, as well as the NUS Institute for Health Innovation & Technology, has invented a new technique called time-resolved assessment of single-cell protein secretion with sequencing (TRAPS-seq) to facilitate the studying of immune cell response at a single-cell level.
"Having progressive snapshots of the dynamic mechanism of immune cells—from responding to a viral infection to eliminating the threat—can accelerate the development of new therapeutic strategies for disease treatment, such as targeting mechanisms that are selective for cancer cells and hence spare healthy cells and reduce side effects," explained Asst. Prof. Cheow. "In addition, a better understanding of how healthy cells communicate can provide valuable insights into the normal cellular processes disrupted in cancer."
This technological breakthrough was reported in Nature Methods on April 10, 2023.
A powerful tool for gathering data on cell functionality
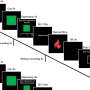
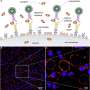
TRAPS-seq employs surface proteins on a cell to anchor the secreted proteins by the cell, which can then be easily analyzed by currently available techniques. The unique advantage of this method over other existing techniques is its capability to track the source cell secreting the proteins and measure the changes in the secretion over time when it responds to a stimulus.
"By modifying the surface proteins on a cell, we can use them as a hook to anchor a secreted protein as soon as it emerges from the cell surface. Using this approach, we can combine with other techniques, such as 10x Genomics barcoding assay, to correlate the cell's functionality directly with its transcription and surface phenotypic profiles," explained Dr. Wu Tongjin, lead author of the paper.
To test the technique's viability, the NUS team used TRAPS-seq to simultaneously monitor three types of secreted cytokine proteins from T-cells, a type of white blood cells linked to our immune defense system to fight against various diseases, by exposing them to a stimulus. The team was able to track the dynamic changes of these secreted proteins over time and observe sub-populations of T-cells that show different dynamic patterns and protein secretion profiles, indicating the potential of using the method to discover new dynamic cellular mechanisms.
Cheow said, "TRAPS-seq provides the opportunity to measure the protein secretion profile of individual cells and resolve their functional differences. Under an abnormal condition, changes to protein secretion would have broad-ranging consequences to our immune system, tissue development and disease progression. The new technique holds great potential as a powerful tool for scientists to discover therapeutic targets for diseases relating to dysregulated protein secretion."
Next steps
Currently, the NUS team can profile three secreted proteins at a time, but the researchers are working towards increasing the number of secreted proteins being studied simultaneously to 10.
The TRAPS-seq assay works for any secreted protein, although immediate applications are likely in immunology profiling. The researchers are optimistic that the extensive catalogs of verified antibody pairs for enzyme-linked immunoassay (ELISA) could be adapted to TRAPS-seq to detect a broad range of secreted proteins for various applications.
More information: Tongjin Wu et al, Time-resolved assessment of single-cell protein secretion by sequencing, Nature Methods (2023). DOI: 10.1038/s41592-023-01841-y