This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
proofread
Researchers identify genes associated with addiction to psychostimulant drugs
Psychostimulant drugs like methamphetamine (METH) and cocaine (COC) affect the brain and nervous system by boosting alertness, attention, and energy levels of the individual. However, their persistent use results in drug addiction, compromising the life of the individual and burdening the health care, social, and legal systems as a consequence.
In order to develop effective prevention and treatment strategies for drug addiction, it is critical to explore how these drugs interact with the nervous system and understand the mechanism underlying the addictive responses of individuals towards psychostimulants.
Interestingly, the gene Period 2 (Per2) has been linked with an increased tendency towards drug abuse. Per2 is associated with our circadian rhythms, our internal clock which regulates our sleep-wake cycle. Previous studies have reported that Per2 knockout (KO) mice, i.e., mice lacking the Per2 gene, show greater addictive responses and withdrawal symptoms towards METH compared to mice with normally functioning or overactive Per2.
Accordingly, Per2 KO mice have been suggested as a potential animal model for understanding the molecular mechanisms underlying susceptibility towards drug abuse. However, its addictive responses to METH and other psychostimulant drugs like cocaine have not been examined as yet.
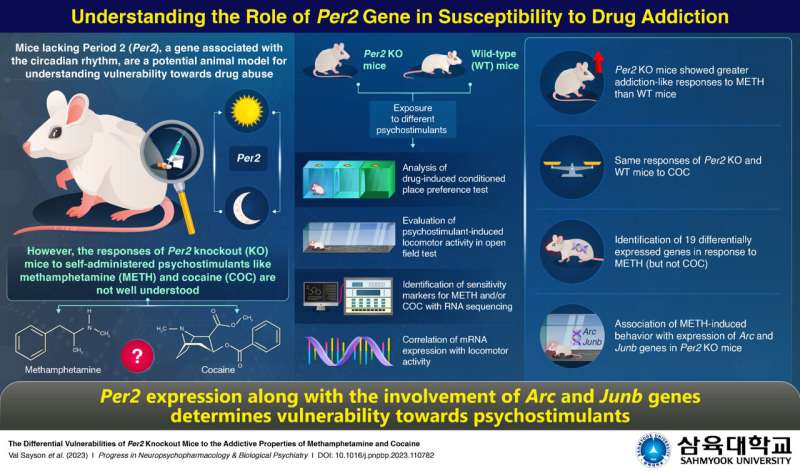
In order to bridge this gap, a group of researchers from Korea and Germany, led by Assistant Professor Hee Jin Kim from Sahmyook University compared how Per2 KO mice and wild-type (WT) mice, i.e., mice with Per2, responded to repeated self-administered doses of METH and COC. Their findings were published in the journal Progress in Neuropsychopharmacology & Biological Psychiatry.
The researchers first investigated the motivational effect of these drugs and their impact on the locomotor activity of the mice using specialized tests. They noted that Per2 KO mice showed a stronger addictive response to METH when compared to the WT mice. However, when it came to drugs like COC, both groups of mice responded in a similar manner.
Highlighting the importance of this observation, Dr. Kim exclaims, "The higher sensitivity of Per2 KO mice to METH rather than COC helped us deduce that different genetic factors were at play behind vulnerability to different drugs."
Probing further, the researchers used a technique that identifies the coding sequence of an RNA–RNA sequencing–to pin down the genes involved in these addictive responses. They identified 19 genes that were only activated in response to repeated doses of METH but not COC. Using existing data, the researchers could correlate these genes with those that are activated in the region of the brain that responds to drug addiction.
Finally, to identify if any of the genes expressed due to METH exposure corresponded with METH-induced behavior, the researchers correlated the mRNA expression levels with the locomotor activity tests in Per2 KO and WT mice. They found that two specific genes, Arc and Junb, were expressed only in Per2 KO mice on METH exposure.
Highlighting their study's findings, Dr. Kim says, "Our findings indicate that Arc and Junb, along with Per2, could be used as markers for susceptibility to METH abuse. If we are able to confirm the role of Arc and Junb expression in vulnerability to drug abuse, they could even be considered as potential targets for treating or preventing drug abuse."
In summary, by unraveling the genetic factors that influence how our brains respond to psychostimulant drugs like METH and COC, researchers can pave the way for more targeted and personalized approaches to tackling drug abuse. In this regard, this study could open doors to the development of more effective diagnostic, preventive and treatment strategies, and potentially save countless lives.
More information: Leandro Val Sayson et al, The differential vulnerabilities of Per2 knockout mice to the addictive properties of methamphetamine and cocaine, Progress in Neuro-Psychopharmacology and Biological Psychiatry (2023). DOI: 10.1016/j.pnpbp.2023.110782















