This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New machine learning techniques boost predictions for virtual drug screening with less data
Scientists using machine learning tools to analyze biomedical data often turn to neural network algorithms, but before these models became popular, another simpler type of machine learning algorithm called kernel methods were commonly used. Kernel methods work by first applying straightforward operations to transform data and then training a simple model on the transformed data.
Now, in a new paper recently published in Nature Communications, researchers at the Eric and Wendy Schmidt Center at the Broad Institute of MIT and Harvard have developed a new way of using kernel methods that could make them more useful for a wider range of applications, such as virtual drug screening.
They came up with the first "transfer learning" techniques for kernel methods that can be successfully applied to large-scale datasets. Transfer learning allows researchers to improve machine learning models by training them on one task in a way that enhances their performance on a second task—without having to spend the time and resources training a new model for each new task.
In their paper, the team showed how their transfer learning framework allowed them to predict which drugs might be most effective in certain cancer cell lines where little data is available. They did this by transferring from cell lines in which many drugs have already been tested.
"Before our paper, there was no transfer learning method for kernel methods that could scale to the large datasets of most interest in the biomedical field and beyond. We've shown for the first time that transfer learning using kernels in these settings is possible and I think that is really exciting," said Caroline Uhler, the senior author on the paper and a Broad core institute member, co-director of the Schmidt Center at Broad, and a professor in the Department of Electrical Engineering and Computer Science as well as the Institute for Data, Systems, and Society at MIT.
The team's key innovation was creatively adapting transfer learning methods used in neural network algorithms so that they can be applied to kernel methods. This advance could find uses in other applications.
"Particularly for health care and biomedical applications, it's very hard to collect a lot of data for every question of interest. When you have very little data for a certain task but a related task has abundant data, this is exactly a setting where our method is effective," said Adityanarayanan Radhakrishnan, a co-first author on the study and a Schmidt Center fellow, who worked on this study while completing his Ph.D. as an Eric and Wendy Schmidt Center Fellow in Uhler's lab at Broad and MIT, and is currently the George F. Carrier Postdoctoral Fellow at Harvard School of Engineering and Applied Sciences.
Transferring knowledge
The research team focused on kernel methods because they found in a previous paper that these performed better than typical neural network models on virtual drug screening tasks. But they wanted to make it possible for researchers to quickly reuse their kernel method algorithms to identify drugs for a wide range of cancer types without having to train a new model for each new type of cancer. They realized that transfer learning techniques are necessary for this, but because existing techniques don't work well for kernel methods, they had to come up with new ones.
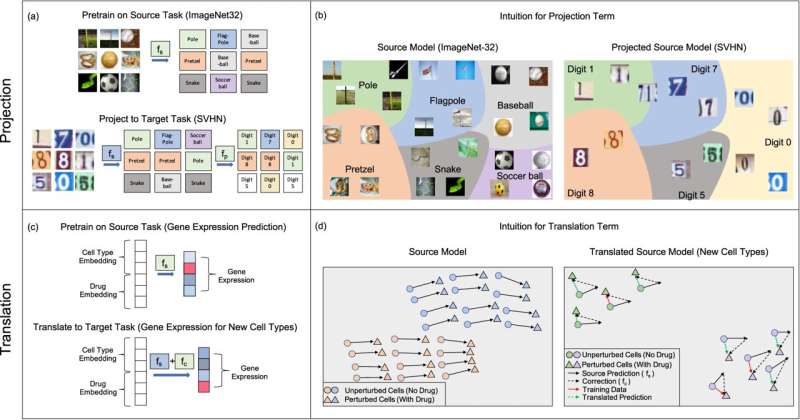
They decided to take inspiration from two transfer learning techniques that work well for neural network models, which they called projection and translation. The team adapted them to work with kernel methods and then tested their approach in a virtual drug screen.
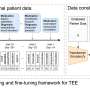
The researchers analyzed performance of their transfer learning algorithms on two massive Broad datasets, one from the Connectivity Map (CMAP) and the other from the Cancer Dependency Map (DepMap). These datasets describe the effects of drugs on cancer cell lines across millions of drug and cell line combinations.
The team trained their kernel method algorithms to predict either the genes expressed by a certain cell type after it was treated with a certain drug (using the CMAP dataset), or the proportion of cancer cells that survived after treatment with the same drug (using the DepMap dataset).
The scientists then applied their projection and translation techniques to their model so that it could complete the second task: to predict the effect of the drug on new cancer cell lines that have much less data. The projection transformation corrects the model's predictions on the second task by recognizing when the prediction errors are falling into categories that can be easily corrected to the right category. And the translation technique fine-tunes the model by applying a correction term that shifts the model's predictions so that it's more accurate on the second task.
The team found that their transfer learning techniques allowed their original kernel method to be successfully "transferred" to the second task, without needing to be retrained. Compared to a new model trained only on the second task, the transfer learning techniques greatly boosted the accuracy of their model in predicting the effect of drugs for new cancer cell lines. And on a common machine learning task where the team trained their kernel method algorithms to recognize images, their approach surprisingly boosted the accuracy by up to 10%.
Moreover, the researchers were also able to pinpoint exactly how much extra data they would need to collect to increase the performance of the model. Uhler said this could be helpful to scientists trying to decide whether it's worthwhile to collect more data in the lab. "That's really quite exciting because you can ask 'how much is it worth for me to have a little bit better performance of my model if I know that we'll need to collect, say, 10% or 20% more data?'" said Uhler.
Beyond drug screening
Two additional advantages of kernel methods are that they provide interpretability as well as a quantification of how uncertain the model is on a given prediction. To take advantage of the interpretability aspect, the research team is working on pinning down the features of a drug that lead their model to predict that it will be effective. In addition, the research team hopes that the uncertainty estimates provided by their kernel approach will be helpful in identifying which new drug and cell line combinations should be screened experimentally for a more effective drug discovery pipeline.
They also have plans to expand their framework to other applications, such as screening cancer genes that tumors heavily depend on for survival and might be targeted with new drugs.
The team adds that their transfer learning approach for kernel methods may also open up other, unexpected applications. Because kernel methods make it easy for scientists to mathematically understand what the model is doing, they can investigate what kinds of biomedical questions will be the best fit to study. "It now gives us a more thorough or deeper understanding of transfer learning and where the power comes from, so that we can analyze which tasks it will actually work for," said Uhler.
More information: Adityanarayanan Radhakrishnan et al, Transfer Learning with Kernel Methods, Nature Communications (2023). DOI: 10.1038/s41467-023-41215-8