This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Research team maps antibiotic-resistant bacteria in Ghana
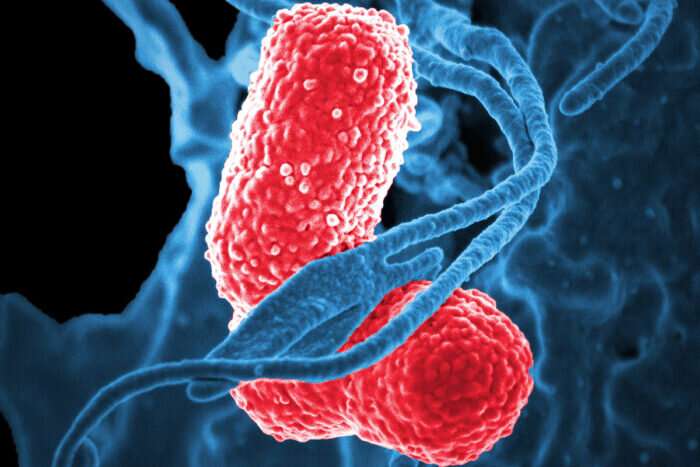
Some strains of heavily antibiotic-resistant bacteria in Ghana are not successful at spreading outside of the hospital, suggesting that control measures can be focused on clinical settings to help curb treatment-resistant infections.
Scientists from the Wellcome Sanger Institute, Oslo University Hospitals, the University for Development Studies in Ghana, and collaborators have used a One Health approach to understand the spread of antibiotic resistance in Klebsiella pneumoniae (K. pneumoniae) bacteria in Ghana. It is a bacterial species that has the ability to cause a wide range of infections.
The study, published in The Lancet Microbe, found that strains of these bacteria that cause typical treatment-resistant infections were only found inside clinical settings, at similar rates to those seen in Italy.
Heavily antibiotic-resistant strains were not found in the environment, or in animals, suggesting that the persistence of these outside the hospitals is currently minimal in this area of Ghana. Researchers point to the clinical use of antibiotics driving this resistance in Klebsiella pneumoniae, and highlight the importance of using genetic data to inform control measures.
Klebsiella pneumoniae, which belongs to the wider Klebsiella genus, is a major human pathogen. It has the ability to cause a wide range of infections, including pneumonia, meningitis, wound and soft tissue infection, urinary tract infections, and is a leading cause of neonatal sepsis.
Antibiotic resistance—otherwise known as multidrug resistance—in K. pneumoniae has been increasing steadily. Some strains of the bacteria are able to produce enzymes known as extended-spectrum beta-lactamases (ESBLs), making them resistant to certain types of antibiotics, including penicillin.
Other strains of K. pneumoniae have become resistant to carbapenems, a broad-spectrum antibiotic used for infections that are not responding to other treatments.
While there has been genomic surveillance of Klebsiella in multiple European countries, there are many countries where the data are still needed.
In this research, the team sequenced 573 Klebsiella samples collected from clinical, environmental, and animal sources in and around the city of Tamale, Ghana. They compared these data to data from previous studies that investigated Klebsiella from Pavia, Italy and Tromsø, Norway.
Among the nearly 600 Klebsiella isolates sequenced, researchers found that K. pneumoniae made up two-thirds of this. The team identified two strains that are resistant to carbapenem, and multiple strains containing ESBL genes, conferring other antibiotic resistance. However, these were only in samples taken from clinical settings. This level of antibiotic resistance was rare in the environmental samples suggesting that these strains are less successful outside of the hospital.
Researchers suggest that the clinical use of antibiotics, such as carbapenems, drove the increase in antibiotic resistance. Without this selective pressure, these resistant strains do not outcompete other, less dangerous forms of the bacteria.
This insight could help inform public health measures focused on reducing the spread of heavily antibiotic-resistant bacteria in hospitals and highlights the importance of genomic surveillance.
Dr. Jessica Calland, first author from Oslo University Hospitals, said, "Being able to map the spread of bacteria that can cause treatment-resistant, and potentially life-threatening, infections is vital in developing methods to help stop this. Our study shows that while antibiotic use has increased the number of resistant strains of Klebsiella pneumoniae, these strains are not as effective at spreading through the wider environment. Identifying which strains outcompete these could be a useful tool in helping to lower the levels of the resistant strains, or inform measures to help curb the spread."
Professor Courage Kosi Setsoafia Saba, co-senior author from the University for Development Studies in Ghana, added, "Treatment-resistant Klebsiella pneumoniae is a growing and ongoing issue in Ghana. However, before this study, we did not have robust genomic surveillance data to understand which strains and types of resistance we were dealing with. Our research shows how important it is to undertake genomic surveillance work in all countries, especially those that are particularly affected by treatment-resistant pathogens, as these may enter the environment because of ineffective treatment of hospital sewage."
Dr. Akosua Karikari, co-senior author from the University for Development Studies in Ghana, stated, "By using the One Health approach, our study takes into account all the different environments in an area, including hospitals, humans and other animals, where bacteria can persist. This allows us to have a more complete view of all the potential pathways in an area, and without this efforts to stop the spread of bacteria could have potential blind spots. Using this approach allowed us to see that multidrug-resistant strains in Ghana were only found in hospitals, validating the extent of misuse of antibiotics in human medicine, and showing where interventions can now be focused."
Professor Jukka Corander, co-senior author from the Wellcome Sanger Institute, noted, "Antibiotic resistance in bacteria is a global public health problem, which impacts every country. Our study highlights this by finding the same genes conferring resistance at similar levels in both Ghana and Italy. Having detailed genomic information on the spread of antibiotic-resistance and the factors that impact this is vital if we are to try to be able to slow and eventually stop this.
"While there have been multiple genomic surveillance studies in European and Western countries, our study works with international collaborators to help inform the situation in a more global setting. Collaboration across borders will be the key to effectively combating antibiotic-resistant bacteria and reducing their negative impact on public health globally."
More information: J K. Calland, K. Haukka, C. K. S. Saba, S W. Kpordze, et al, Population structure and antimicrobial resistance among Klebsiella isolates sampled from human, animal, and environmental sources in Ghana: a cross-sectional genomic One Health study, The Lancet Microbe (2023). DOI: 10.1016/S2666-5247(23)00208-2