This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Translating the language of brain cells to figure out what goes wrong in diseases of the brain
Despite all our cells sharing the same DNA, there are thousands of different cell types in the human brain, each with a unique structure and function. One longstanding problem in neuroscience is determining how genes are switched on and off to form the mosaic of different cell types within the brain. Scientists from University of California San Diego School of Medicine have published two new studies that bring us closer to solving this mystery.
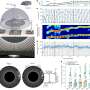
The researchers analyzed more than 2.3 million individual brain cells from mice to create a comprehensive map of the mouse brain and used artificial intelligence to help predict what stretches of DNA are used to determine a brain cell's type. The researchers also looked at the brains of humans and primates to study the evolution of the processes cells use to turn genes on and off. The findings are published December 14, 2023 in a special edition of Nature.
"A cell's DNA is like its language," said senior author Bing Ren, Ph.D., professor at UC San Diego School of Medicine. "Just like there are certain root words that many languages share, there are certain genes and gene expression patterns that are conserved across different species. Learning to understand and interpret the brain's molecular language can help us learn more about how the brain works in general and about what happens to the brain in neuropsychiatric conditions."
The two new papers are part of a package of 10 studies describing the first complete cell type atlas of a mammalian brain, led by researchers at UC San Diego, the Salk Institute for Biological Studies, the Allen Institute for Brain Science and other institutions.
The research is part of the National Institutes of Health's Brain Research Through Advancing Innovative Neurotechnologies Initiative, or the BRAIN Initiative, which launched in 2014 to deepen our understanding of the inner workings of the human mind and improve how we treat, prevent and cure disorders of the brain.
"This work is helping us establish a baseline understanding of what the brain is like at the cellular level," said Ren. "This will make it possible to draw comparisons between our baseline and brains with neurological and psychiatric disorders. Studying the brain this way could help us discover new therapeutic approaches for these conditions."
One of the most ambitious projects under the Brain Initiative is the Cell Census Network (BICNN), which seeks to describe human brain cells in unprecedented molecular detail, classifying them into more precise subtypes, pinpointing their locations in the brain and tracking how cellular features change over a lifetime.
Earlier this year, Ren and other scientists from the BICCN published a first-of-its kind atlas of the human brain, which identified more than a hundred types of brain cell. Their new atlas of the mouse brain complements this work and expands upon it by drawing comparisons between the brains of different species.
For example, by comparing the brains of mice with those of humans and nonhuman primates, the researchers found that cell-type-specific patterns of gene expression evolve much more rapidly than patterns that are shared across cell types. This could help explain why there are so many different cell types in the brain.
"Humans have evolved over millions of years, and much of that evolutionary history is shared with other animals," said Joseph Ecker, Ph.D., a professor at the Salk Institute for Biological Studies who co-led one of the new studies with Ren.
"Data from humans alone is never going to be enough to tell us everything we want to know about how the brain works. By filling in these gaps with other mammalian species, we can continue to answer those questions and improve the machine-learning models we use by providing them more data."
While the BRAIN Initiative and BICCN are still very much ongoing projects, some insights are already proving relevant to human diseases. For example, the researchers found that many of the genetic programs that determine cell type were in parts of the genome that have already been implicated in human diseases, such multiple sclerosis, anorexia nervosa and tobacco use disorder. This could help shed light on how neuropsychiatric disorders affect the brain.
"The brain isn't homogenous, and diseases don't affect all parts of the brain equally," said Ren. "Insights from this research and the BRAIN initiative as a whole are helping us better understand what types of cells are affected in specific diseases. We hope this will pave the way for more precise, targeted therapies that can heal diseased cells without affecting the rest of the brain."
More information: Single-cell analysis of chromatin accessibility in the adult mouse brain, Nature (2023). DOI: 10.1038/s41586-023-06824-9. www.nature.com/articles/s41586-023-06824-9
Conserved and divergent gene regulatory programs of the mammalian neocortex, Nature (2023). DOI: 10.1038/s41586-023-06819-6. www.nature.com/articles/s41586-023-06819-6