This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Study reveals new opportunities to develop cancer treatments

Researchers at Baylor College of Medicine and collaborating institutions have uncovered new potential therapeutic targets for cancer and new insights into existing cancer drug targets, expanding the breadth of possibilities for treating this disease.
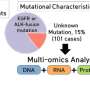
Using a comprehensive approach that included integrating proteomics, genomics and epigenomics data from 10 cancer types, the team identified protein and small protein or peptide targets in cancer tissues and validated many of them experimentally as promising candidates for therapeutic strategies. The study appeared in Cell.
"Experience has shown that targeted therapies, cancer treatments directed at specific proteins in cancer cells, hold promise for achieving more effective clinical results than traditional radiotherapy and chemotherapy," said co-corresponding author Dr. Bing Zhang, professor of molecular and human genetics and part of the Lester and Sue Smith Breast Center at Baylor.
"Although there is progress identifying potential vulnerabilities of specific cancer types, fewer than 200 proteins are targeted by FDA-approved cancer drugs. In this study we significantly expanded the list of potential therapeutic targets by analyzing data from more than 1,000 tissue samples spanning 10 cancer types."
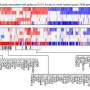
The researchers applied computational tools to integrate proteogenomic data comprising genome-wide information on DNA, RNA and proteins that was generated by the Clinical Proteomic Tumor Analysis Consortium (CPTAC) from prospectively collected samples of treatment-naive primary tumors, many with matched normal adjacent tissues for comparison.
The team integrated the CPTAC dataset with other public datasets to investigate the similarities and differences among gene and protein alterations found in diverse tumor types to illuminate protein targets for cancer therapy.
"Our goal was to better understand the characteristics of known drug targets. We also hoped to identify new targets that could lead to new drug developments," said Zhang, a McNair scholar and member of Baylor's Dan L Duncan Comprehensive Cancer Center.
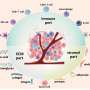
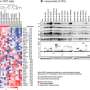
The team applied the data integration approach to systematically identify proteins and genes that are important for cancer growth and development. For instance, proteins that are overexpressed or hyperactive in cancer tissues but not in normal counterparts, and loss of tumor suppressor genes, which can create dependencies on other proteins that could then be therapeutically targeted. They also searched for tumor antigens, including neoantigens—cancer-specific peptides derived from gene mutations in tumors.
"Our study revealed new opportunities for repurposing drugs currently approved for other conditions," Zhang said. "For example, we show that an antifungal drug can also reduce growth of several cancer types, supporting further exploration of the anti-cancer value of this drug."
The researchers also identified potential protein targets currently without a drug—some are enzymes called kinases and others are cell surface proteins. "These findings open opportunities for drug development, including small-molecule drugs or drug-antibody conjugates," Zhang said.
Furthermore, computational identification of several tumor-associated proteins shared among different cancer types was followed by experimental confirmation of their importance for cancer in cells grown in the lab and in animal models, validating these proteins as potential therapeutic targets worthy of more study.
"I am very excited that we have created a comprehensive resource of protein targets, significantly expanding the therapeutic landscape by identifying many new candidates and covering various therapeutic modalities. And we have made our findings publicly available at https://targets.linkedomics.org," Zhang said. "We hope that this resource will pave the way to repurposing currently-available drugs and developing new therapies for cancer treatment."
More information: Pan-cancer proteogenomics expands the landscape of therapeutic targets, Cell (2024). DOI: 10.1016/j.cell.2024.05.039. www.cell.com/cell/fulltext/S0092-8674(24)00583-X