Sequencing cancer mutations - there's an app for that
Using precise information about an individual’s genetic makeup is becoming increasingly routine for developing tailored treatments for breast, lung, colon and other cancers. But techniques used to identify meaningful gene mutations depend on analyzing sequences of both normal and mutant DNA in tumor samples, a process that can yield ambiguous results. Now, a team of Johns Hopkins researchers says it has developed an easy-to-use online computer software application that can clear up any confusion faster and cheaper than other methods currently used to do the job.
The application, called “Pyromaker,” is available free-of-charge at pyromaker.pathology.jhmi.edu/ , and a related tutorial will be posted there soon. The software generates simulated pyrograms, which are readouts from a gene sequencing technique known as pyrosequencing.
Most pyrograms correspond precisely to a person’s unique mutation or set of mutations, but some mutations can be more difficult to interpret than others, the Johns Hopkins researchers say.
“Pyromaker’s value is in rapidly sorting through each of several simulated pyrograms, until there is a clear match with the actual tumor pyrogram,” says James R. Eshleman, M.D., Ph.D., a professor in the departments of Pathology and Oncology at Johns Hopkins. “Pyromaker enables us to do in minutes, essentially at no cost, what otherwise would take days of further, expensive tests.”
Eshleman led the team that developed Pyromaker, the software code for which was written by Johns Hopkins pathology resident Matthew T. Olson. A report on the application, with demonstrations of Pyromaker’s ability to resolve sequencing ambiguities, was published online in the Journal of Molecular Diagnostics.
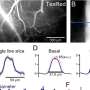
Swedish researchers invented pyrosequencing in the mid-1990s as an alternative to the traditional method of gene sequencing. Pyrosequencing is an automated process in which solutions of the four DNA-building nucleotides, or “bases” (G, C, T, A) are added, one at a time, to single strands of DNA from sampled cells, in order to build up a complementary matching strand. A match with the next available base causes a chemical reaction with a special enzyme, which in turn emits a light pulse, resulting in a “peak” on the pyrogram – effectively indicating the next base in the DNA sequence.
Pyrosequencing works on shorter stretches of DNA than does the traditional method, known as “Sanger,” named for Frederick Sanger who invented the process. But pyrosequencing is also more sensitive in registering the presence of mutant DNA in a tumor sample, which is a mix of tumor and normal cells. That sensitivity makes it very useful for tumor sequencing, says Eshleman, because the mutant genes that drive a tumor’s abnormal growth typically are less prevalent in a tumor sample, compared with normal versions of those genes.
Because a tumor pyrogram is an overlay of both healthy and mutant DNA, identifying the correct sequence may be difficult and further studies to sort it all out can delay diagnosis and add significantly to costs, he says.
To develop the software, Eshleman and his team first confirmed that simulated pyrograms generated by Pyromaker matched actual pyrograms of tumor DNA containing well-known, common cancer-driving mutations in the genes KRAS, BRAF, GNAS and p53. Then, they focused on the example of KRAS, an “oncogene” that helps drive many tumor types, to show that nearly all one- and two-base mutations in KRAS have distinguishable pyrograms.
In a few cases, the researchers found that the pyrograms of two different mutations appear identical under certain sequencing conditions. With Pyromaker, they were able to show that pyrograms of two such mutations would become readily distinguishable if they were re-sequenced with a different order of added bases.
Finally, the researchers applied Pyromaker to ambiguous KRAS pyrograms from two colorectal cancer patients, and compared the results with tests using Sanger sequencing and other relatively time-consuming methods to resolve the ambiguities. The comparison showed that Pyromaker could resolve the ambiguities more quickly.
Eshleman notes that Pyromaker also may be used with a new sequencing technology known as ion semiconductor sequencing, which detects a change in hydrogen ions instead of a light pulse.
The software was written by Guoli Chen and Matthew T. Olson, both from Eshleman’s laboratory in the Department of Pathology at Johns Hopkins. Other scientists who contributed to the research include Alan O’Neill, Alexis Norris, Katie Beierl, Shuko Harada, Marija Debeljak, Keila Rivera-Roman, Samantha Finley, Amanda Stafford, Christopher David Gocke, and Ming-Tseh Lin, all from Johns Hopkins.
Contributions to funding the research came from the Sol Goldman Pancreatic Cancer Research Center, the Michael Rolfe Foundation, the Dick Knox Foundation, and the National Institutes of Health (CA130938).