Go with your gut: Research sheds light on how microbes can interact with drugs

Scientists are already working to develop treatments that can be tailored to an individual's genetics, but what about tailoring treatments based on the genetics of the trillions of microbes that live in a person's gut?
The idea might not be as far-fetched as it sounds, said Peter Turnbaugh, a Bauer Fellow at Harvard's Faculty of Arts and Sciences (FAS) Center for Systems Biology.
In a recent paper in Cell, Turnbaugh and co-authors Corinne Ferrier Maurice and Henry Joseph Haiser, both postdoctoral fellows at the Center for Systems Biology, show that, as drugs are administered, the activity of human gut microbes can change dramatically. Understanding how those changes affect drugs could one day help researchers to design drugs that work more effectively and antibiotics that more specifically target pathogens.
"The big question is: To what extent do the benefits and side effects of different types of drugs depend on the microbes in our gut?" Turnbaugh said. "For decades, we have known they can play some role. The microbes inhabiting our gut are able to change the structure of drugs in ways that can contribute to toxicity, or to activate or inactivate compounds. In almost all cases, however, we don't know the particular microbes that are responsible, the genes they may be using, or the factors that promote or inhibit these activities."
Previously, Maurice and Haiser could examine how the microbes in the gut reacted to various treatments. However, they wanted to know exactly which microbes were there, and whether all were equally active.
To get at that question, they turned to a technique borrowed from aquatic microbial ecology.
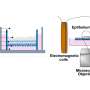
They first marked cells with a series of three stains designed to highlight whether and how severely cells were damaged, and how active the cells were, based on the amount of DNA and RNA they contained. Using a flow cytometer, a device that uses lasers to count and sort cells precisely, they were able to determine how many of each type of cell were present in the samples.
"Our initial view suggests that the gut microbiota is quite active relative to other environments, and there's also a substantial percentage—around 30 percent—of damaged cells," Turnbaugh said. "We also found that both the active and damaged groups were primarily made up of Firmicutes, one of the two major groups of bacteria in the gut. That suggests that the Firmicutes may be more highly active than other members of our gut microbial community."
Armed with that data, researchers used next-generation sequencing to study how the gene expression of the bacteria changed as six drugs and eight antibiotics were administered.
"We know that at least some members of the community are able to change these drugs. Our hope was that finding changes in gene expression would give us a clue as to who is responsible and what genes they might be using," Turnbaugh said. "We were able to identify a variety of changes in gene expression, many of them consistent with the known biochemical changes to each compound, providing a starting point for more mechanistic studies."
For the various antibiotics they tested, Turnbaugh said, colleagues were surprised to see different responses between individuals and for each individual on different days.
"The main goal of antibiotics is to eliminate pathogenic bacteria, but we're learning that there are many side effects that these drugs have on the microbes that are normally found in the gut, which may have negative consequences," he said. "There has been a great deal of emphasis on personalized medicine in recent years, and the standard way of thinking about that is understanding the human genome and trying to predict how a given drug will react inside your body. I think this paper emphasizes that it may be equally important to think about how your particular gut microbiota will interact with a given drug."
This story is published courtesy of the Harvard Gazette, Harvard University's official newspaper. For additional university news, visit Harvard.edu.