Drug-resistant MRSA bacteria: Here to stay in both hospital, community
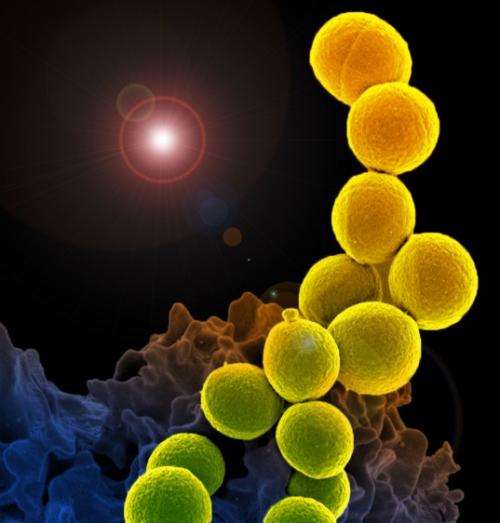
(Medical Xpress)—The drug-resistant bacteria known as MRSA, once confined to hospitals but now widespread in communities, will likely continue to exist in both settings as separate strains, according to a new study.
The prediction that both strains will coexist is reassuring because previous projections indicated that the more invasive and fast-growing community strains would overtake and eliminate hospital strains, possibly posing a threat to public health.
Researchers at Princeton University used mathematical models to explore what will happen to community and hospital MRSA strains, which differ genetically. Originally MRSA, which is short for methicillin-resistant Staphylococcus aureus, was confined to hospitals. However, community-associated strains emerged in the past decade and can spread widely from person to person in schools, athletic facilities and homes.
Both community and hospital strains cause diseases ranging from skin and soft-tissue infections to pneumonia and septicemia. Hospital MRSA is resistant to numerous antibiotics and is very difficult to treat, while community MRSA is resistant to fewer antibiotics.
The new study found that these differences in antibiotic resistance, combined with more aggressive antibiotic usage patterns in hospitals versus the community setting, over time will permit hospital strains to survive despite the competition from community strains. Hospital-based antibiotic usage is likely to successfully treat patients infected with community strains, preventing the newcomer strains from spreading to new patients and gaining the foothold they need to out-compete the hospital strains.
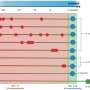
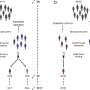
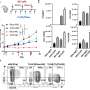
The researchers made their predictions by using mathematical models of MRSA transmission that take into account data on drug-usage, resistance profiles, person-to-person contact, and patient age.
More information: Kouyos R., Klein E. & Grenfell B. (2013). Hospital-Community Interactions Foster Coexistence between Methicillin-Resistant Strains of Staphylococcus aureus. PLoS Pathogens, 9 (2) e1003134. PMID: 23468619: www.plospathogens.org/article/info%3Adoi%2F10.1371%2Fjournal.ppat.1003134