Methylation linked to metabolic disease
(Medical Xpress)—In the first in-depth analysis of DNA methylation in fat, a process that affects the regulation of genes, researchers have linked regions of methylation to metabolic traits such as high body mass index (BMI) and obesity.
Charting DNA methylation is vital to understanding genetic regions associated with disease. These chemical tags form part of the epigenome: the chemical and structural changes to DNA that affect the function of a person's genome. The team examined the genomes of fat cells taken from twins and found that DNA methylation patterns are very likely to be passed down from parents to their children.
Twin studies can help to tease out to what extent variation in DNA methylation is genetic and how much is due to environmental factors encountered in a lifetime. For twins that are genetically identical from birth, differences in disease development are likely to be due to environmental influences.
DNA methylation is an epigenetic change but is often under genetic control. The most common form of DNA methylation involves the addition of a molecule, known as a methyl group, to the DNA base cysteine. This process is vital to the regulation of many cellular processes such as embryonic development, transcription, and chromatin structure. Consistent with these important roles, a growing number of human diseases have been associated with aberrant DNA methylation.
"We are exploring epigenetic variation in people, trying to understand the interplay between gene expression and DNA methylation and how they impact disease," says Professor Panos Deloukas, from the Wellcome Trust Sanger Institute and Queen Mary University London. "Our research has uncovered many links between methylation and genetic regions in fat tissue associated with metabolic diseases."
The team surveyed more than 450,000 sites on the genome from fat tissue samples from 648 twins, the largest twin study of its kind. They found that, in apparently healthy people, there is little variation in methylation patterns. Variability is suppressed in regions that are important for gene regulation such as promoters.
When the team integrated genetic variation and DNA methylation they found 28% of sites to be under genetic control. Overlapping this information with the variants known to regulate gene expression showed that only 6 per cent of these variants regulated both gene expression and DNA methylation.
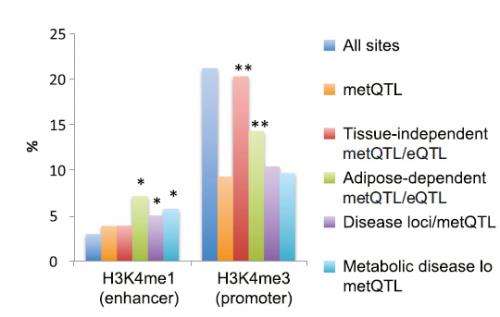
The team integrated the information about DNA methylation with other genetic data from the NIH RoadMap Epigenomics Mapping Consortium, and all known regions in the genome associated with common disease. They found many of the variants that regulate DNA methylation overlap with metabolic-trait or disease loci in regulatory elements known as gene enhancers. The strongest effects were seen for high-density lipoprotein cholesterol and BMI.
"We know that DNA methylation plays a role in gene regulation and disease susceptibility," says Elin Grundberg, lead author from the Wellcome Trust Sanger Institute and McGill University, Montreal, Canada. "Through our research we are beginning to unravel the frequency, location and function of DNA methylation variation."
This study provides functional information for genetic regions associated with metabolic disease. The team did further analysis on a variant known to be associated with increased BMI, close to the ADCY3 gene. They found that this variant also regulates DNA methylation in the same region.
This study has greatly enhanced the understanding of the link between methylation and disease in specific cell types. The next step for the team is to explore further the effects of methylation on other tissue types such as blood and skin cells and start mapping shared versus cell-restricted sites of DNA methylation.
More information: "Global Analysis of DNA Methylation Variation in Adipose Tissue from Twins Reveals Links to Disease-Associated Variants in Distal Regulatory Elements," Elin Grundberg, Eshwar Meduri, Johanna K. Sandling, sa K. Hedman, Sarah Keildson, Alfonso Buil, Stephan Busche, Wei Yuan, James Nisbet, Magdalena Sekowska, Alicja Wilk, Amy Barrett, Kerrin S. Small, Bing Ge, Maxime Caron, So-Youn Shin, the Multiple Tissue Human Expression Resource Consortium, Mark Lathrop, Emmanouil T. Dermitzakis, Mark I. McCarthy, Timothy D. Spector2 Jordana T. Bell, and Panos Deloukas. American Journal of Human Genetics 2013, DOI: 10.1016/j.ajhg.2013.10.004