New understanding of chlamydial disease: Novel simultaneous RNA-Seq analysis tracks host/pathogen interactions
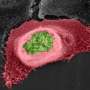
Investigators at the Institute for Genome Sciences at the University of Maryland School of Medicine have developed a new technique that can track the activity of a disease-causing microbe and the host cell response to that pathogen simultaneously. Using the new method to examine Chlamydia trachomatis infection, the study team observed how the response of the infected cell contributes to one of the hallmark outcomes of chlamydial disease—tissue scarring. Their findings appear in the December 4 issue of PLOS One.
Chlamydia trachomatis is an intracellular, disease-causing bacterium responsible for the most common human sexually transmitted infections (STIs) and infectious blindness (trachoma) globally. Sexually transmitted chlamydial infections are often asymptomatic, and cause tissue damage and scarring. For example, chlamydial-induced scar tissue within the fallopian tubes can block the tubal opening and lead to infertility. In trachoma, chlamydial scarring of the tissue lining the inside of the eyelids leads to eyelash inversion and direct abrasion of the cornea by the eyelashes, ultimately resulting in the cornea turning opaque.
The central paradox of chlamydial infection is that human immune and cellular responses to the infection contribute to disease. An interdisciplinary team of genome scientists and bioinformatics experts were able to develop an innovative method using new RNA-Seq technology to reveal the complex interplay between invading bacterial pathogens and their host mammalian cells.
"Next generation sequencing technology has advanced so that we can now apply these sophisticated analytical tools to complex bacterial infections of human cells, even at very early points of infections. These early events often set the stage for disease to occur much later," says Garry Myers, Ph.D., Assistant Professor of Microbiology and Immunology at the Institute for Genome Sciences (IGS) at the University of Maryland School of Medicine and senior author on the paper. "We found that the response to chlamydial infection is rapid and dramatic, and observed many novel host cell transcriptional reactions to the infection. In particular, we were able to identify abnormal early expression of host cell genes known to induce scarring, and expression of numerous genes that encode the building blocks of fibrotic scars," said Myers. "It seems that a series of dominos start to fall as soon as Chlamydia infect a cell. Depending on how the individual reacts to that infection, these host responses induce a series of positive feedback loops that ultimately amplify production of the disease-causing scars over time."
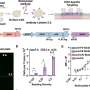
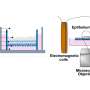
In this study, the scientists used depletion of both bacteria and human rRNA to enrich bacterial and human RNA from infected cells for simultaneous sequencing. This next generation deep sequencing method distinguished chlamydial and host expression, yielding a detailed view of both host and pathogen transcription, particularly in the poorly characterized early stages of infection.
"With this simultaneous RNA-Seq approach, we were able to examine how Chlamydia and the infected host cell responded to each other. This gives us significant insight into scarring in chlamydial disease," said Myers. "The RNA-Seq approach pioneered here is also applicable to any bacteria that infect human cells."
"The simultaneous RNA-Seq method developed in this project is likely to be widely used in host/pathogen studies," says Claire M. Fraser, Ph.D., director of the Institute for Genome Sciences and the principal investigator of the National Institutes of Health-funded Genomic Sequencing Center for Infectious Diseases, which conducted this work. "This is a spectacular example of how technology development related to study of a specific pathogen can be leveraged more broadly."
"The marriage between the fields of genomics and bioinformatics provides new tools for basic research scientists to understand how harmful microbes cause disease," says E. Albert Reece, M.D., Ph.D., M.B.A., Vice President for Medical Affairs at the University of Maryland and the John Z. and Akiko K. Bowers Distinguished Professor and Dean of the University of Maryland School of Medicine. "The ability to examine the interactions between the host cells and the pathogen gives investigators a more complete picture of infection and could uncover new therapeutic targets."