Faster fish thanks to nMLF neurons
As we walk along a street, we can stroll at a leisurely pace, walk quickly, or run. The various leg movements needed to do this are controlled by special neuron bundles in the spinal cord. It is not quite clear how these central pattern generators know how quickly the legs are to be moved. An international team working with scientists from Harvard University and the Max Planck Institute of Neurobiology in Martinsried has now discovered individual neurons in the brain of zebrafish larvae that control the animals' swimming speed. Human movements are also controlled by central pattern generators. The results represent an important step in gaining a better understanding of how rhythmic movements are modulated.
As young children, we learn how to place one foot in front of the other at a steady pace. Once this has been learned, small bundles of neurons in the spinal cord - the central pattern generators (CPG) - ensure that this sequence happens almost automatically: we do not need to think about when and how far away we should place our foot down when we take each step. Once they are operational, the CPG neurons do not need any further stimulus to transmit their impulses. But how are these cells stimulated, and how does the brain tell them how quickly the legs need to be moved?
Ruben Portugues and his colleagues have studied zebrafish larvae to investigate how the brain and the CPGs are connected. The animals use various methods to increase their speed: they can beat their tails for longer periods of time, move the tail to and fro more vigorously, reduce the time between periods of tail movements such that these periods called bouts happen more frequently or switch to a completely different movement rhythm or gait - like a horse that changes from a trot to a gallop.
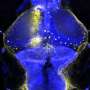
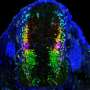
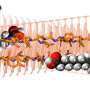
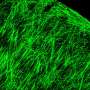
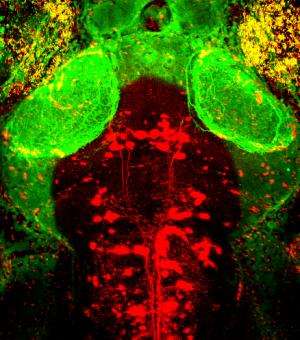
To understand how the brain triggers these various types of swimming movements, the neurobiologists concentrated on a group of around 20 neurons which send out their extensions from the midbrain to the spinal cord. The scientists already knew that the cells in this nMLF region are active during swimming. They were now able to show that stimulating these cells triggered swimming movements. As the researchers now report in the journal Neuron, the cells in the central pattern generator receive the initial stimulus for a movement from neurons in the nMLF region. They also discovered that it is almost impossible for the fish to regulate their swimming speed if four particular nMLF cells are switched off.
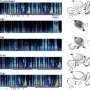
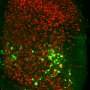
Calcium-sensitive dyes can be used to image neuronal activity. As zebrafish larvae are transparent, the scientists were able to observe the activity of individual nMLF cells directly through the microscope. "It was especially exciting when the animals changed their speed," reports Ruben Portugues, who was recently appointed Leader of a Research Group at the Max Planck Institute of Neurobiology. "We had actually expected that more nMLF cells would simply be activated simultaneously to enable the fish to swim faster."
Instead, the scientists discovered that neurons which were already active became even more active when swimming faster. "We don't yet know the details of how a higher level of activity leads to faster movements," says Portugues. However, the scientists can demonstrate that individual nMLF cells, known as MeLR cells, control the length of swimming periods and MeLc cells, as they are known, control the frequency to the tail beating. To date, scientists were aware of the nMLF region and its cells, but nobody knew what they control or how they do it. "Now that we have found the transmission, as it were, for the swimming movements, the next question to be answered is how and where the brain decides what gear it wants to engage," says Ruben Portugues, summing up the next challenge.
More information: Kristen E. Severi, Ruben Portugues, João C. Marques, Donald M. O'Malley, Michael B. Orger, Florian Engert, "Neural Control and Modulation of Swimming Speed in the Larval Zebrafish," Neuron, Available online 24 July 2014, ISSN 0896-6273, DOI: 10.1016/j.neuron.2014.06.032.