Discovery of new genes involved in the parasitism of cells by the tubercle bacillus
A European-Asian collaboration of scientists have identified ten virulence genes of the tubercle bacillus. The inactivation of these genes lessens the pathogenic effects of the bacillus. This discovery, published in the journal PLoS Pathogens, will make it possible in particular to propose original therapeutic strategies and to test novel anti-tuberculosis vaccine candidates. In order to obtain these results in only two weeks, the researchers developed a new screening technique. This innovative, rapid and efficient method could be easily transposed to other intracellular pathogens.
Despite existing medicines and BCG vaccination, tuberculosis continues to wreak havoc: every year, it kills nearly 2 million people throughout the world. This disease is caused by bacteria of the mycobacterium family, which include Mycobacterium tuberculosis, the agent of TB in humans. Its virulence, in other words its pathogenicity, depends on its capacity to propagate within the host cell. This pathogenic agent is in fact capable of escaping the defenses of the host, which it infects through parasitism of the macrophages, the immune system cells normally involved in the ingestion and destruction of microbes. Following inhalation, Mycobacterium tuberculosis enters the lungs where it is ingested by the alveolar macrophages and lies in an intracellular compartment known as a "phagosome".
Normally, the function of a phagosome is to destroy the bodies ingested by the macrophage through acidification. However, instead of being killed by the cell, the tubercle bacillus multiplies within it by blocking the acidification of the phagosome. The genes of the bacillus involved in this process were poorly understood until now. Identifying them was one of the key objectives of the European-Asian collaboration coordinated by Olivier Neyrolles, CNRS researcher at the Institut de Pharmacologie et de Biologie Structurale and Priscille Brodin, Inserm researcher (Institut Pasteur Korea / Inserm).
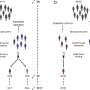
To achieve these results, the researchers firstly designed a new screening method based on the selection, by a robot, of a cellular phenotype characterized by a microscopy image. Using this innovative technique, samples corresponding to the requisite criterion can be automatically and visually identified. This procedure thus makes it possible to gain considerable time in identifying the microbial entities involved in the parasitism of cells.
This novel and highly efficient method was then applied to a particularly virulent strain of Mycobacterium tuberculosis. This enabled more than 11,000 mutants of the tubercle bacillus to be screened in just a few weeks. The robot was then programmed to detect whether the “acidification” function was active or not. This allowed the researchers to isolate the mutants unable to block acidification of the phagosome and subsequently destroyed by the macrophages. The corresponding mutations were identified by genetic engineering and ten genes, mostly unknown, involved in the parasitism of the macrophage were characterized.
Most of these genes code for the synthesis of products secreted by the bacteria: proteins and lipids, whose precise function in the parasitism of the cells by Mycobacterium tuberculosis still needs to be elucidated. In addition, these molecules could constitute prime targets for new antibiotics. Finally, these results suggest that the isolated mutants, the in vivo virulence of which is lessened in some cases, could be promising candidates in the development of new vaccines to replace the existing BCG vaccine.
More information: High content phenotypic cell-based visual screen identifies Mycobacterium tuberculosis acyltrehalose-containing glycolipids involved in phagosome remodeling. Priscille Brodin, et al. PLoS Pathogens, 9 September 2010