Swapping 'dance partners' in the brain is key to learning

(PhysOrg.com) -- Researchers collected brain imaging data from people performing a motor task, and then analysed this data using new computational techniques. They found evidence that the 'flexibility' of a person's brain - how much different areas of the brain link up in different combinations; essentially 'swapping partners' - can be used to predict how fast someone will learn.
An international team - from Oxford University, UC Santa Barbara, and UNC Chapel Hill - publish a report of their research in this week's Proceedings of the National Academy of Sciences.
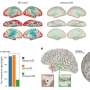
The team ran an experiment over 3 sessions in which 18 volunteers had to push a series of buttons, rather like a sequence of notes on a piano keyboard, as fast as possible. They then divided functional MRI images of each volunteer's brain into 112 different regions and analysed how these different areas partnered up - that is, were active together - whilst they performed the task.
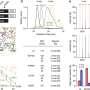
The researchers found that it was possible to predict faster learning of new sequences in the later sessions using the 'flexibility' of the brain - how often different areas of the brain 'swapped partners' by changing the regions that were jointly active - in the earlier sessions. They found that people with more 'flexible' brains, whose brain regions swapped partners more often, were faster at learning the motor tasks than those whose brains exhibited less frequent partner-swapping.
"It's the first time that anyone has defined this concept of 'flexibility' in the brain: how brain regions 'light up' together in different combinations. We've been able to show that how much these areas 'swap partners' in one session can predict how fast people will perform a task in a later session,' said Dr. Mason Porter of Oxford University's Mathematical Institute, an author of the report. 'It suggests that in order to learn the networks of our brains have to be flexible."
The new study uses computational methods developed to analyse what the researchers call 'multilayer' networks, in which each 'layer' might represent a network at one snapshot in time or a different set of connections between the same set of regions or individuals. These layers are combined into a larger mathematical object, which can contain a potentially huge amount of data and is difficult to analyse. Previous methods could only deal with each layer separately, and it was necessary to compare the results obtained from different layers using ad hoc tools (if it was possible at all) and the new method overcomes this key challenge.
Dr. Porter said: "With this technique, we are treating the brain like a social network in which each of the 112 regions is analogous to a person, then looking at how they interact with each other over time. One of the reasons the study is small is that, even taking fMRI data from a small number of individuals over three sessions, this turns out to be an immense computing task: this project included 10,000 days of computing (CPU) time to process, though we're now working on making our computations more efficient to be able to handle larger amounts of data from experiments."
The team also had to develop new ways of randomising different combinations of brain regions to ensure that the connections they were seeing were part of the brain's 'dance' and not just something that could be attributed to chance. They are now in the process of analysing a study in which participants took part in sessions spread out over 30 days.
Dr. Porter added: 'The hope is that with a longer study, we might be able to use these techniques to pinpoint where during a process what we think of as 'learning' is actually occurring. This would be a great step forward in understanding the human brain, which of course is one of the most complex of all networks.'
More information: PNAS paper