Scientists solve mystery of nerve disease genes
For several years, scientists have been pondering a question about a genetic disease called Charcot-Marie-Tooth (CMT) disease type 2D: how can different types of mutations, spread out across a gene, produce the same condition?
Now, a team of scientists at The Scripps Research Institute may have found the answer. By studying a gene called GARS, which is mutated in individuals with the disease, the team found that all the mutations have one thing in common: they cause the tightly coiled three-dimensional shape of the resulting protein to shift open.
The more open configuration creates a space for other proteins to bind, causing havoc. "That is the basis for potential disease-causing interactions," said Scripps Research Associate Professor Xiang-Lei Yang, senior author of the study, "but also for potential therapeutic intervention." It is possible that scientists could develop drugs to fit into the open area, blocking its access to other proteins.
The findings, to be published this week in the online issue of Proceedings of the National Academies of the Sciences (PNAS) help scientists explain how CMT type 2D occurs. The results may also have implications for other diseases, such as amyotrophic lateral sclerosis (ALS), also known as Lou Gehrig's disease.
A Puzzling Disease
Named after the three physicians who first reported the disease in 1886, CMT is the most common inherited neurological disorder, estimated to affect one in 2,500 people. Although it is not fatal, CMT causes progressive weakness and wasting of muscles in the feet, legs, hands, and forearms by striking down the nerves that reach down into these muscles.
There are five different forms of CMT, and each is broken down in various subtypes depending on the gene responsible. CMT type 2D, which is caused by mutations in the GARS gene, is inherited in an autosomal dominant manner—this means that a person needs to inherit only one faulty copy of the GARS gene from one parent to get the disease.
The GARS gene holds the instructions for producing an enzyme called glycyl-tRNA synthase (GlyRS), which is vital to the process by which amino acids, the building blocks of proteins, are attached to one another during protein synthesis. So far, scientists have found 11 different kinds of mutations in the GARS gene that cause CMT type 2D. Some of the mutations affect the protein-building function of the GlyRS enzyme, but others don't, suggesting that a change in enzyme activity is not what causes CMT.
Thus, scientists have been trying to find a common feature shared by all GlyRS mutants that might explain the disease mechanism.
Structures in Solution
To find this common feature, Yang and colleagues turned to the three-dimensional structure of GlyRS. In 2007, they published the X-ray crystal structure of the wild-type protein and one of the mutants, but the structures were not all that different from each other (PNAS 104:11239-44, 2007).
"When we compared the wild type with the mutant, we did not find dramatic conformational change," said Weiwei He, a Scripps Research graduate student who joined Yang's lab and took on the project around the time of the 2007 paper's publication. "We thought maybe it was because of the crystal packing."
X-ray crystallography requires obtaining high amounts of a protein and packing the protein molecules into crystals that are bombarded with X-rays to determine the positions of all the atoms. This preparation, Yang and He reasoned, may mask any subtle conformation changes in a protein. "That is why we thought to look at the structure in solution, under more physiological conditions," said He.
In collaboration with Hui-Min Zhang, a postdoctoral fellow in the laboratory of Alan Marshall at the National High Magnetic Field Laboratory at Florida State University, Yang and He used a method to obtain the structures of five different GlyRS mutants and the wild-type protein in solution. This method, called hydrogen–deuterium exchange, gives information about which parts of a protein are in touch with a solution—in other words, which parts lie on the outside rather than being buried deep inside its three-dimensional shape. In collaboration with Min Guo, assistant professor at Scripps Florida, a second method, called small angle X-ray scattering, was used to measure changes in the overall shape of the protein structure in solution.
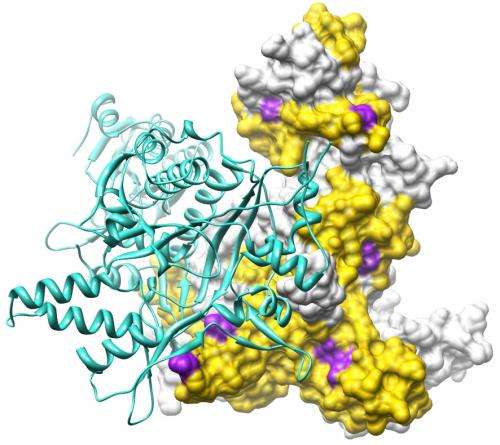
Together, the results revealed that every GARS mutation studied causes a structural opening in the resulting mutant protein. By superimposing the information obtained from the structures in solution to the high-resolution X-ray crystal structure of the wild-type GlyRS protein, Yang, He, Zhang and colleagues were able to determine that the structural opening maps to a common region of the GlyRS protein in all the mutants.
By becoming more open to potential new partners, the GlyRS mutants may gain a new function that is toxic to nerve cells. That would explain why CMT type 2D is inherited in a dominant fashion.
Exciting Implications
The team is now looking for proteins that might bind to this region and already have leads on several candidates. As exciting as this work is, says He, "this paper is just the start of the story; we have more story coming, and it's getting more and more exciting."
This disease mechanism may also apply to other conditions. "Some proteins may be relatively unstable and can be easily triggered into another conformation by different types of mutations," explained Yang. "This example deals with the GlyRS protein, but the general idea can be applied to many other mutation-induced human diseases."
One example may be ALS. Some inherited forms of ALS are caused in a gene that encodes the enzyme copper-zinc superoxide dismutase (commonly called SOD1). To date, more than 100 different mutations in the SOD1 gene have been linked to inherited ALS.
In addition to He, Zhang, Guo, Marshall, and Yang, Yeeting E. Chong of Scripps Research is a co-author of the paper, "Dispersed disease-causing neomorphic mutations on a single protein promote the same localized conformational opening."