Scientists ID compounds that target amyloid fibrils in Alzheimer's, other brain diseases
UCLA chemists and molecular biologists have for the first time used a "structure-based" approach to drug design to identify compounds with the potential to delay or treat Alzheimer's disease, and possibly Parkinson's, Lou Gehrig's disease and other degenerative disorders.
All of these diseases are marked by harmful, elongated, rope-like structures known as amyloid fibrils, linked protein molecules that form in the brains of patients.
Structure-based drug design, in which the physical structure of a targeted protein is used to help identify compounds that will interact with it, has already been used to generate therapeutic agents for a number of infectious and metabolic diseases.
The UCLA researchers, led by David Eisenberg, director of the UCLA–Department of Energy Institute of Genomics and Proteomics and a Howard Hughes Medical Institute investigator, report the first application of this technique in the search for molecular compounds that bind to and inhibit the activity of the amyloid-beta protein responsible for forming dangerous plaques in the brain of patients with Alzheimer's and other degenerative diseases.
In addition to Eisenberg, who is also a professor of chemistry, biochemistry and biological chemistry and a member of UCLA's California NanoSystems Institute, the team included lead author Lin Jiang, a UCLA postdoctoral scholar in Eisenberg's laboratory and Howard Hughes Medical Institute researcher, and other UCLA faculty.
The research was published July 16 in eLife, a new open-access science journal backed by the Howard Hughes Medical Institute, the Max Planck Society and the Wellcome Trust.
A number of non-structural screening attempts have been made to identify natural and synthetic compounds that might prevent the aggregation and toxicity of amyloid fibrils. Such studies have revealed that polyphenols, naturally occurring compounds found in green tea and in the spice turmeric, can inhibit the formation of amyloid fibrils. In addition, several dyes have been found to reduce amyloid's toxic effects, although significant side effects prevent them from being used as drugs.
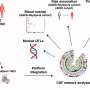
Armed with a precise knowledge of the atomic structure of the amyloid-beta protein, Jiang, Eisenberg and colleagues conducted a computational screening of 18,000 compounds in search of those most likely to bind tightly and effectively to the protein.
Those compounds that showed the strongest potential for binding were then tested for their efficacy in blocking the aggregation of amyloid-beta and for their ability to protect mammalian cells grown in culture from the protein's toxic effects, which in the past has proved very difficult. Ultimately, the researchers identified eight compounds and three compound derivatives that had a significant effect.
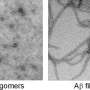
While these compounds did not reduce the amount of protein aggregates, they were found to reduce the protein's toxicity and to increase the stability of amyloid fibrils—a finding that lends further evidence to the theory that smaller assemblies of amyloid-beta known as oligomers, and not the fibrils themselves, are the toxic agents responsible for Alzheimer's symptoms.
The researchers hypothesize that by binding snugly to the protein, the compounds they identified may be preventing these smaller oligomers from breaking free of the amyloid-beta fibrils, thus keeping toxicity in check.
An estimated 5 million patients in the U.S. suffer from Alzheimer's disease, the most common form of dementia. Alzheimer's health care costs in have been estimated at $178 billion per year, including the value of unpaid care for patients provided by nearly 10 million family members and friends.
In addition to uncovering compounds with therapeutic potential for Alzheimer's disease, this research presents a new approach for identifying proteins that bind to amyloid fibrils—an approach that could have broad applications for treating many diseases.